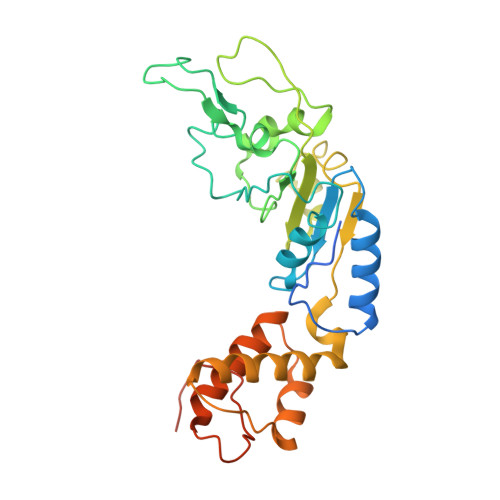
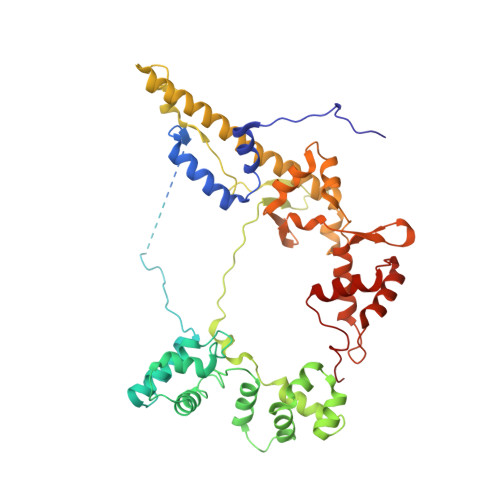
Mechanisms of DNA opening revealed in AAA+ transcription complex structures.
Ye, F., Gao, F., Liu, X., Buck, M., Zhang, X.(2022) Sci Adv 8: eadd3479-eadd3479
- PubMed: 36542713 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.add3479
- Primary Citation Related Structures:
7QV9, 7QWP, 7QXI - PubMed Abstract:
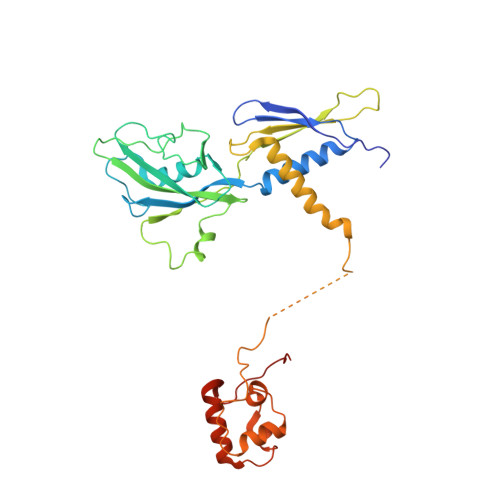
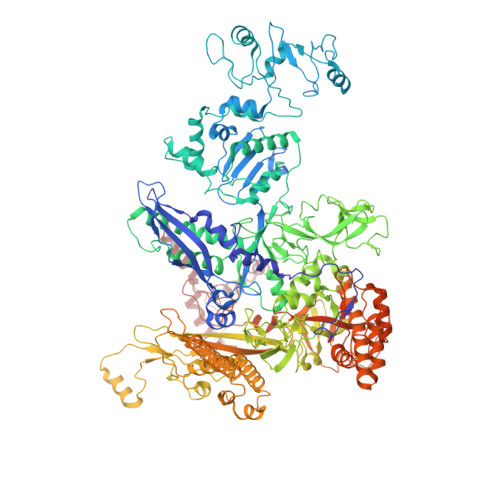
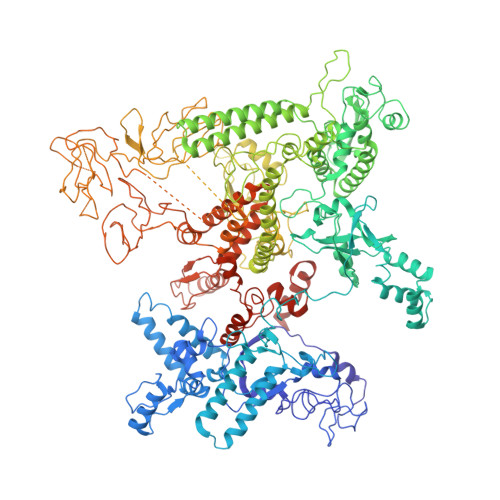
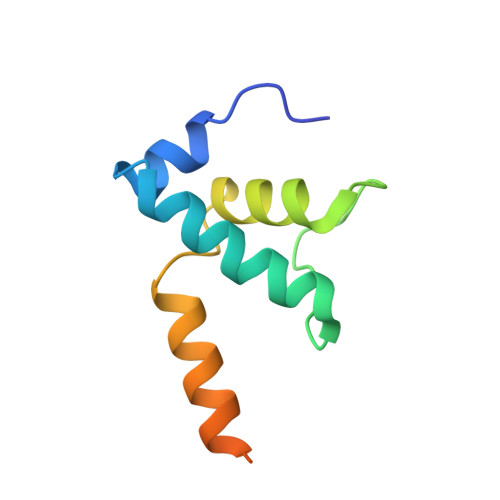
Gene transcription is carried out by RNA polymerase (RNAP) and requires the conversion of the initial closed promoter complex, where DNA is double stranded, to a transcription-competent open promoter complex, where DNA is opened up. In bacteria, RNAP relies on σ factors for its promoter specificities. Using a special form of sigma factor (σ 54 ), which forms a stable closed complex and requires its activator that belongs to the AAA+ ATPases (ATPases associated with diverse cellular activities), we obtained cryo-electron microscopy structures of transcription initiation complexes that reveal a previously unidentified process of DNA melting opening. The σ 54 amino terminus threads through the locally opened up DNA and then becomes enclosed by the AAA+ hexameric ring in the activator-bound intermediate complex. Our structures suggest how ATP hydrolysis by the AAA+ activator could remove the σ 54 inhibition while helping to open up DNA, using σ 54 amino-terminal peptide as a pry bar.
- Section of Structural and Synthetic Biology, Department of Infectious Disease, Faculty of Medicine, Imperial College London, South Kensington SW7 2AZ, UK.
Organizational Affiliation: