High-affinity peptides developed against calprotectin and their application as synthetic ligands in diagnostic assays.
Diaz-Perlas, C., Ricken, B., Farrera-Soler, L., Guschin, D., Pojer, F., Lau, K., Gerhold, C.B., Heinis, C.(2023) Nat Commun 14: 2774-2774
- PubMed: 37198182 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38075-7
- Primary Citation Related Structures:
7QUV - PubMed Abstract:
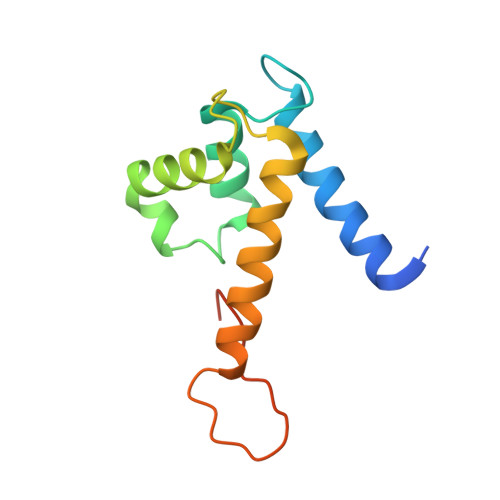
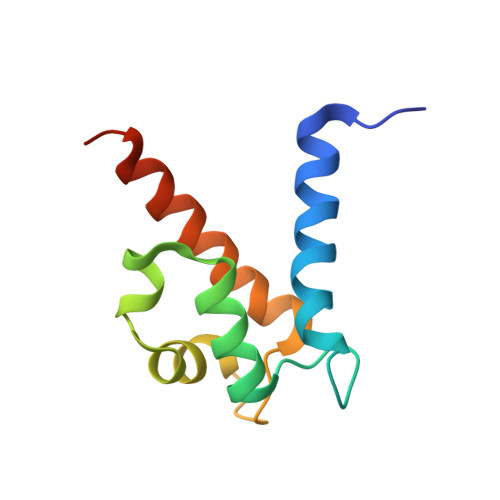
Common inflammatory disorders such as ulcerative colitis and Crohn's disease are non-invasively diagnosed or monitored by the biomarker calprotectin. However, current quantitative tests for calprotectin are antibody-based and vary depending on the type of antibody and assay used. Additionally, the binding epitopes of applied antibodies are not characterized by structures and for most antibodies it is unclear if they detect calprotectin dimer, tetramer, or both. Herein, we develop calprotectin ligands based on peptides, that offer advantages such as homogenous chemical composition, heat-stability, site-directed immobilization, and chemical synthesis at high purity and at low cost. By screening a 100-billion peptide phage display library against calprotectin, we identified a high-affinity peptide (K d = 26 ± 3 nM) that binds to a large surface region (951 Å 2 ) as shown by X-ray structure analysis. The peptide uniquely binds the calprotectin tetramer, which enabled robust and sensitive quantification of a defined species of calprotectin by ELISA and lateral flow assays in patient samples, and thus offers an ideal affinity reagent for next-generation inflammatory disease diagnostic assays.
- Institute of Chemical Sciences and Engineering, School of Basic Sciences, École Polytechnique Fédérale de Lausanne (EPFL), CH-1015, Lausanne, Switzerland.
Organizational Affiliation: