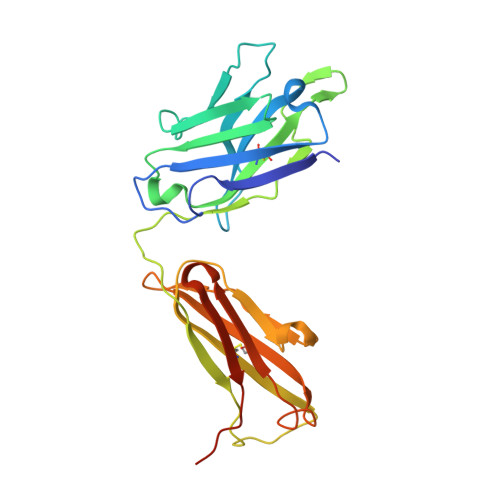
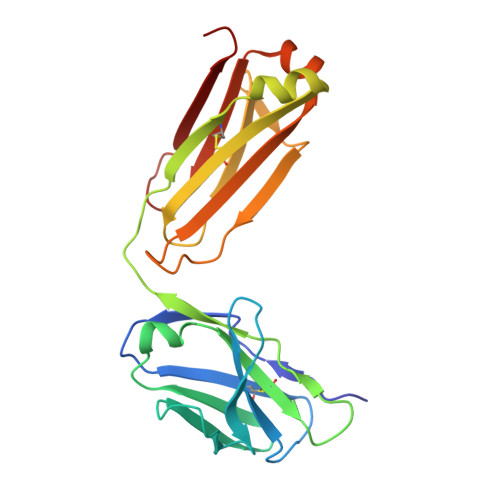
Contrasting Modes of New World Arenavirus Neutralization by Immunization-Elicited Monoclonal Antibodies.
Ng, W.M., Sahin, M., Krumm, S.A., Seow, J., Zeltina, A., Harlos, K., Paesen, G.C., Pinschewer, D.D., Doores, K.J., Bowden, T.A.(2022) mBio 13: e0265021-e0265021
- PubMed: 35315691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.02650-21
- Primary Citation Related Structures:
7QU1, 7QU2 - PubMed Abstract:
Transmission of the New World hemorrhagic fever arenaviruses Junín virus (JUNV) and Machupo virus (MACV) to humans is facilitated, in part, by the interaction between the arenavirus GP1 glycoprotein and the human transferrin receptor 1 (hTfR1). We utilize a mouse model of live-attenuated immunization with envelope exchange viruses to isolate neutralizing monoclonal antibodies (NAbs) specific to JUNV GP1 and MACV GP1. Structures of two NAbs, termed JUN1 and MAC1, demonstrate that they neutralize through disruption of hTfR1 recognition. JUN1 utilizes a binding mode common to all characterized infection- and vaccine-elicited JUNV-specific NAbs, which involves mimicking hTfR1 binding through the insertion of a tyrosine into the receptor-binding site. In contrast, MAC1 undergoes a tyrosine-mediated mode of antigen recognition distinct from that used by the reported anti-JUNV NAbs and the only other characterized anti-MACV NAb. These data reveal the varied modes of GP1-specific recognition among New World arenaviruses by the antibody-mediated immune response. IMPORTANCE The GP1 subcomponent of the New World arenavirus GP is a primary target of the neutralizing antibody response, which has been shown to be effective in the prevention and treatment of infection. Here, we characterize the structural basis of the antibody-mediated immune response that arises from immunization of mice against Junín virus and Machupo virus, two rodent-borne zoonotic New World arenaviruses. We isolate a panel of GP1-specific monoclonal antibodies that recognize overlapping epitopes and exhibit neutralizing behavior, in vitro . Structural characterization of two of these antibodies indicates that antibody recognition likely interferes with GP1-mediated recognition of the transferrin receptor 1. These data provide molecular-level detail for a key region of vulnerability on the New World arenavirus surface and a blueprint for therapeutic antibody development.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxfordgrid.4991.5, Oxford, United Kingdom.
Organizational Affiliation: