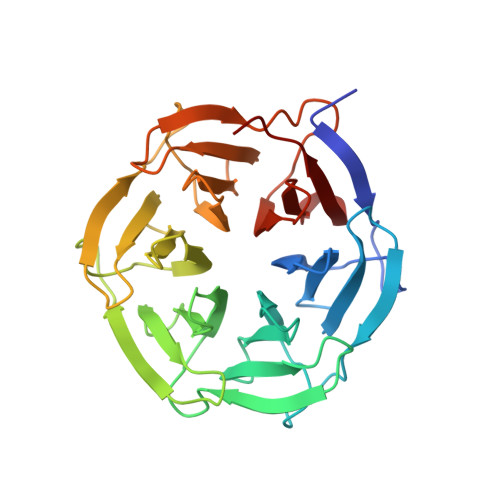
Comparative structural analyses of the NHL domains from the human E3 ligase TRIM-NHL family.
Chaikuad, A., Zhubi, R., Tredup, C., Knapp, S.(2022) IUCrJ 9: 720-727
- PubMed: 36381143 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252522008582
- Primary Citation Related Structures:
7QRV, 7QRW, 7QRX - PubMed Abstract:
Tripartite motif (TRIM) proteins constitute one of the largest subfamilies of the RING-type E3 ubiquitin ligases that play a role in diverse processes from homeostasis and immune response to viral restriction. While TRIM proteins typically harbor an N-terminal RING finger, a B-box and a coiled-coil domain, a high degree of diversity lies in their C termini that contain diverse protein interaction modules, most of which, both structures and their roles in intermolecular interactions, remain unknown. Here, high-resolution crystal structures of the NHL domains of three of the four human TRIM-NHL proteins, namely TRIM2, TRIM3 and TRIM71, are presented. Comparative structural analyses revealed that, despite sharing an evolutionarily conserved six-bladed β-propeller architecture, the low sequence identities resulted in distinct properties of these interaction domains at their putative binding sites for macromolecules. Interestingly, residues lining the binding cavities represent a hotspot for genetic mutations linked to several diseases. Thus, high sequence diversity within the conserved NHL domains might be essential for differentiating binding partners among TRIM-NHL proteins.
- Institute for Pharmaceutical Chemistry, Johann Wolfgang Goethe-University, Max-von-Laue-Strasse 9, D-60438 Frankfurt am Main, Germany.
Organizational Affiliation: