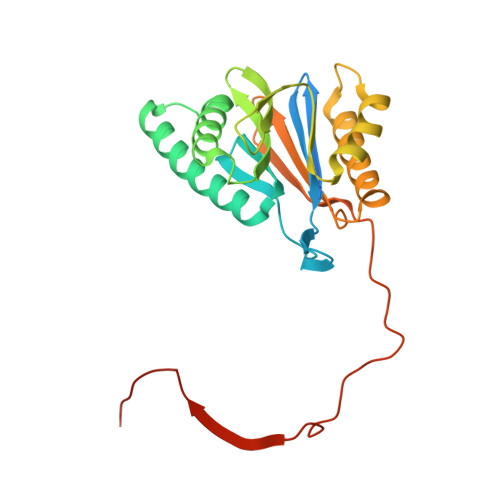
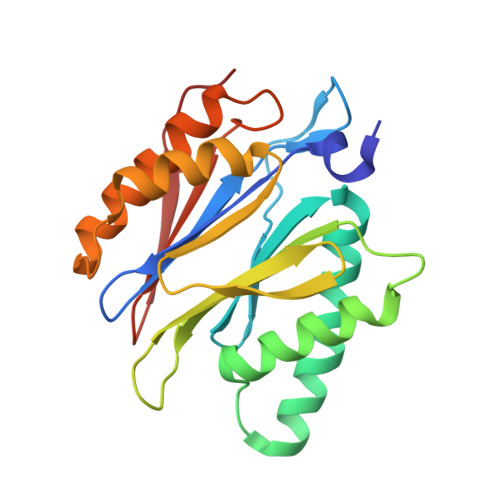
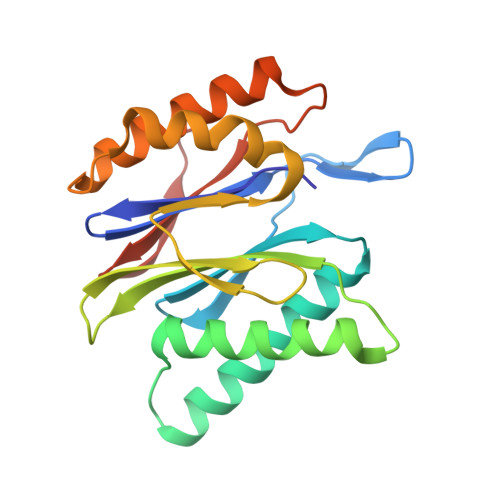
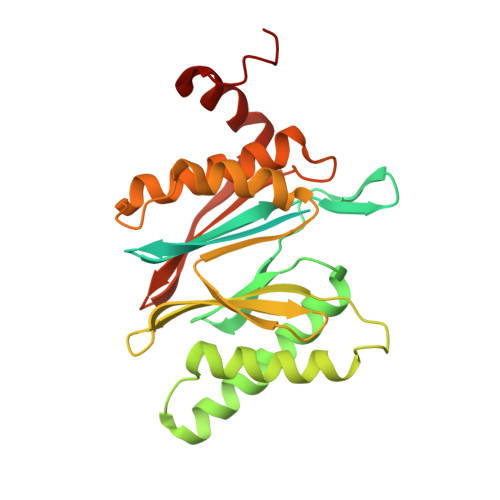
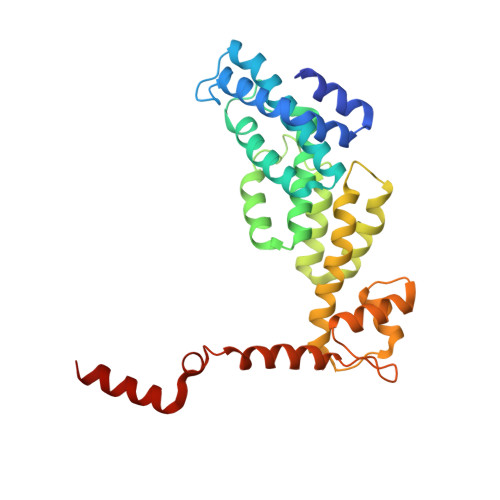
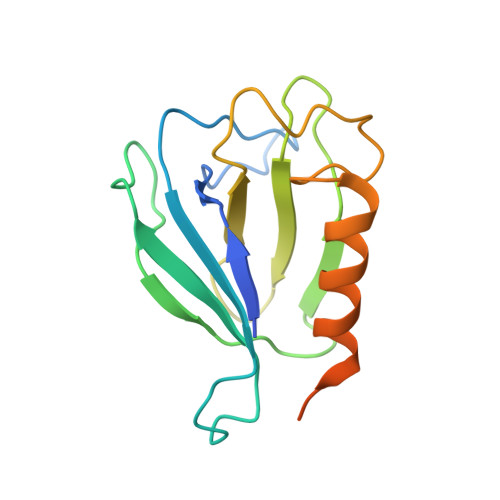
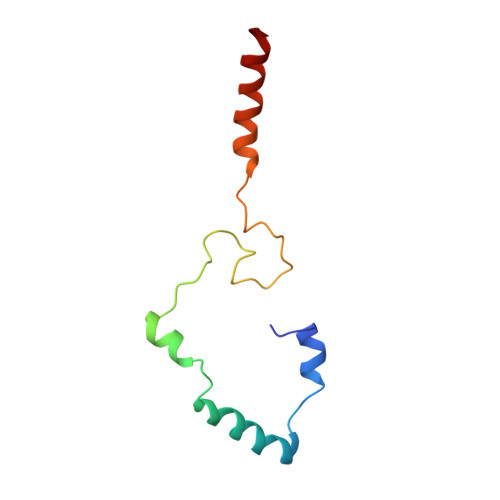
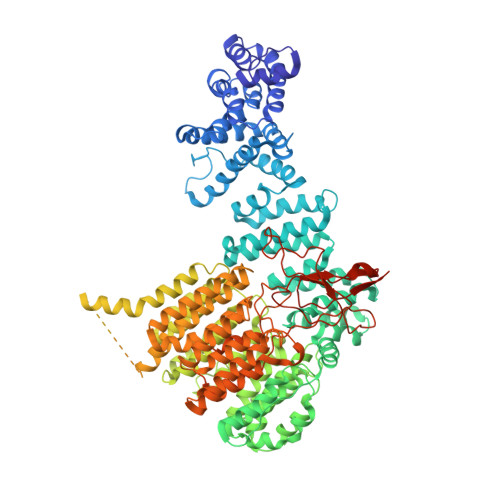
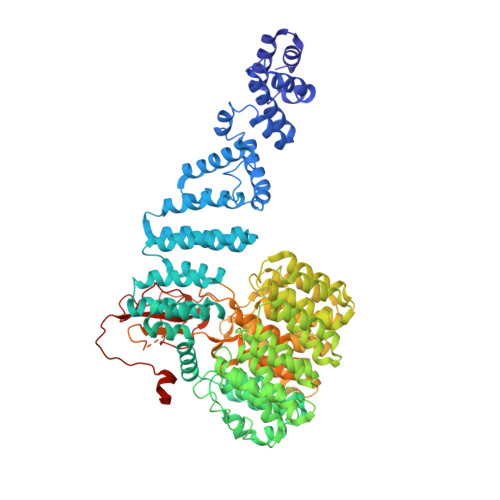
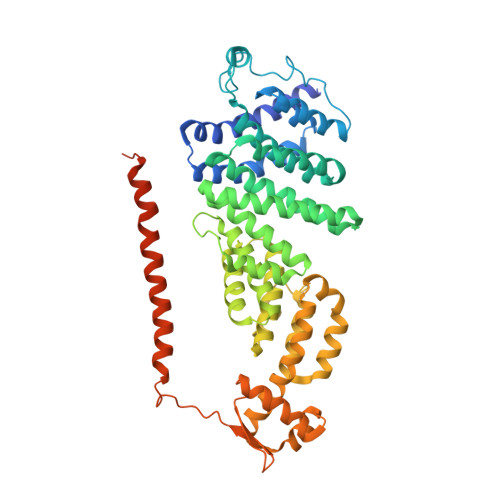
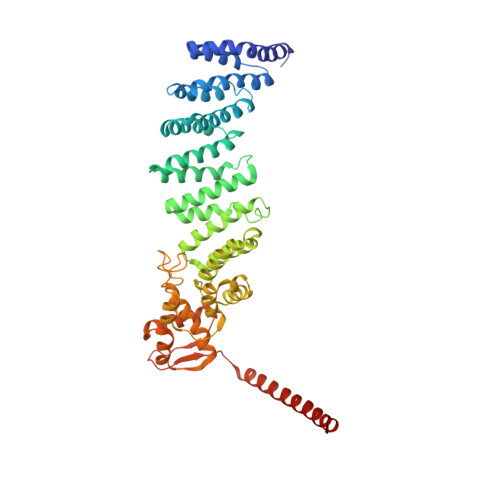
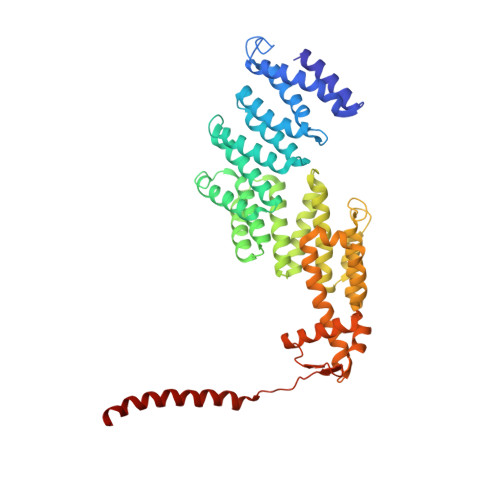
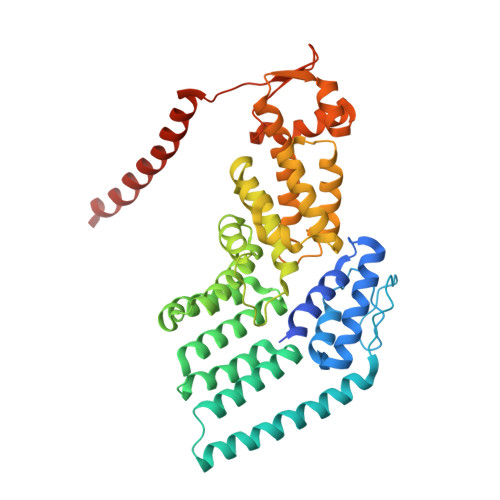
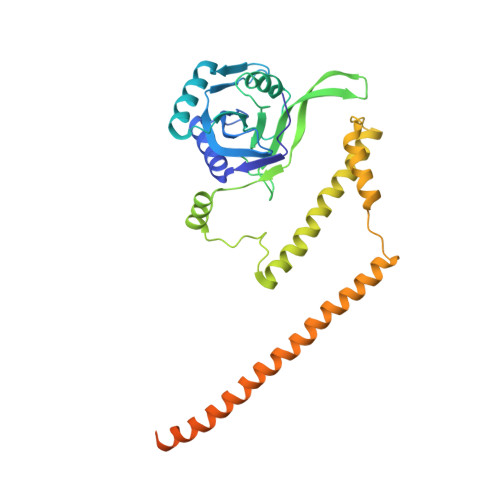
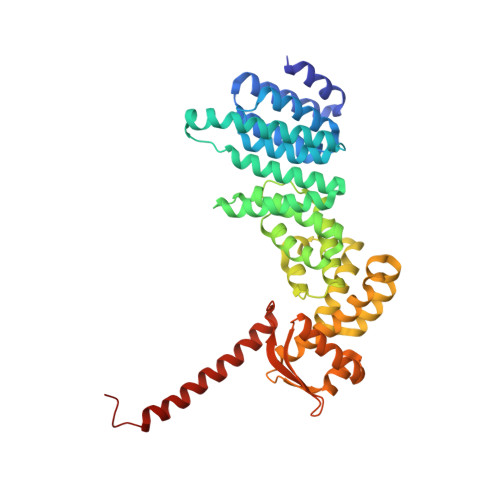
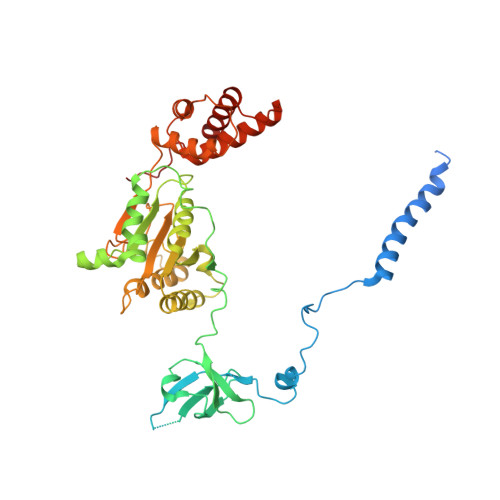
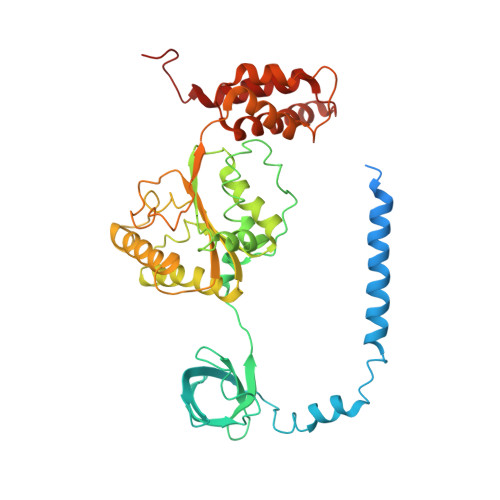
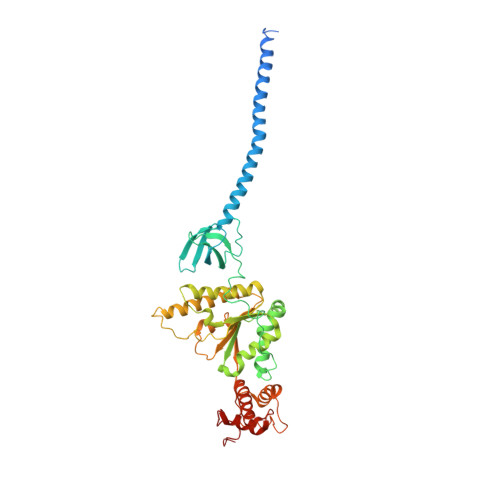
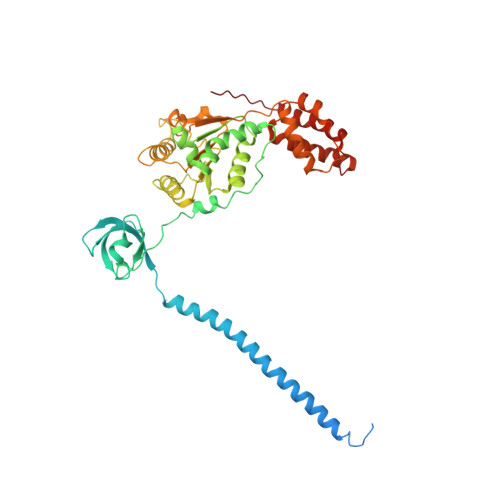
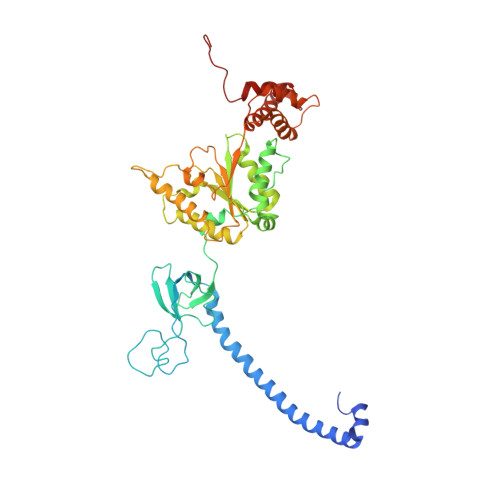
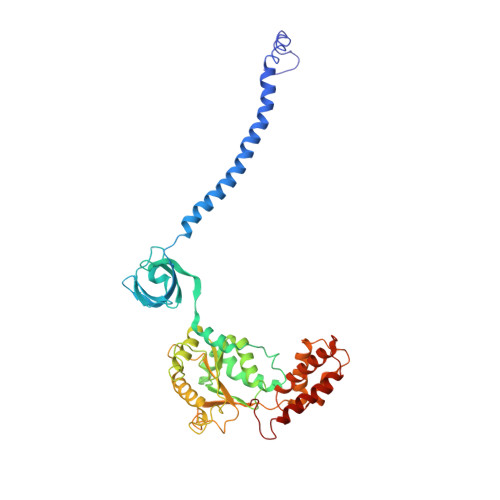
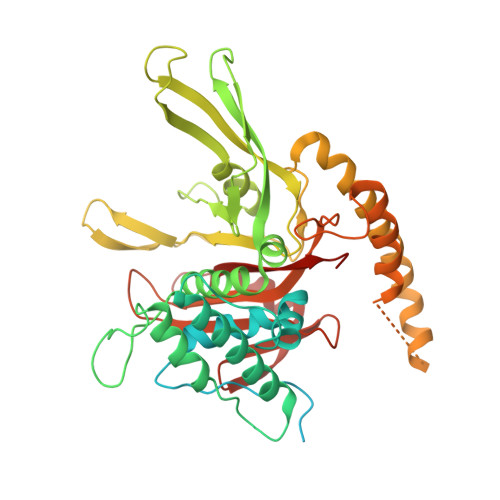
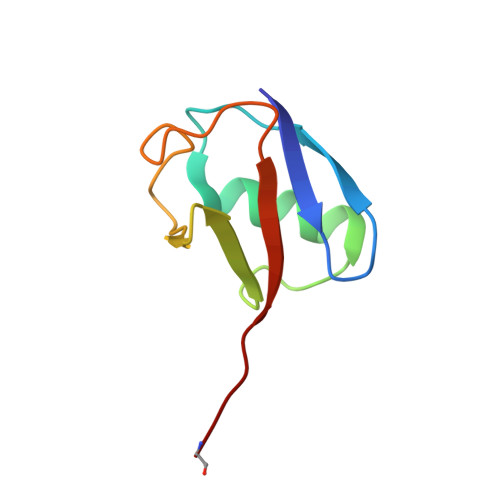
Allosteric control of Ubp6 and the proteasome via a bidirectional switch.
Hung, K.Y.S., Klumpe, S., Eisele, M.R., Elsasser, S., Tian, G., Sun, S., Moroco, J.A., Cheng, T.C., Joshi, T., Seibel, T., Van Dalen, D., Feng, X.H., Lu, Y., Ovaa, H., Engen, J.R., Lee, B.H., Rudack, T., Sakata, E., Finley, D.(2022) Nat Commun 13: 838-838
- PubMed: 35149681 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28186-y
- Primary Citation Related Structures:
7QO3, 7QO5, 7QO6 - PubMed Abstract:
The proteasome recognizes ubiquitinated proteins and can also edit ubiquitin marks, allowing substrates to be rejected based on ubiquitin chain topology. In yeast, editing is mediated by deubiquitinating enzyme Ubp6. The proteasome activates Ubp6, whereas Ubp6 inhibits the proteasome through deubiquitination and a noncatalytic effect. Here, we report cryo-EM structures of the proteasome bound to Ubp6, based on which we identify mutants in Ubp6 and proteasome subunit Rpt1 that abrogate Ubp6 activation. The Ubp6 mutations define a conserved region that we term the ILR element. The ILR is found within the BL1 loop, which obstructs the catalytic groove in free Ubp6. Rpt1-ILR interaction opens the groove by rearranging not only BL1 but also a previously undescribed network of three interconnected active-site-blocking loops. Ubp6 activation and noncatalytic proteasome inhibition are linked in that they are eliminated by the same mutations. Ubp6 and ubiquitin together drive proteasomes into a unique conformation associated with proteasome inhibition. Thus, a multicomponent allosteric switch exerts simultaneous control over both Ubp6 and the proteasome.
- Department of Cell Biology, Harvard Medical School, Boston, MA, 02115, USA.
Organizational Affiliation: