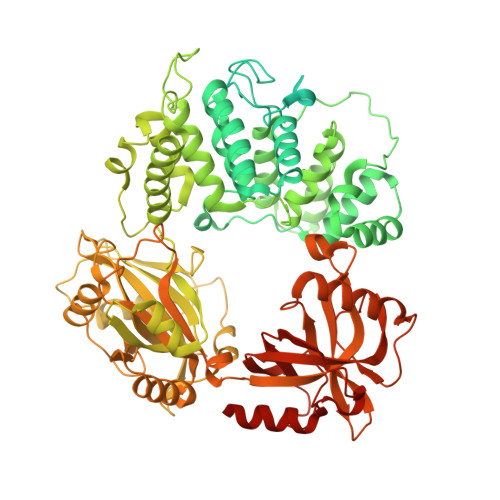
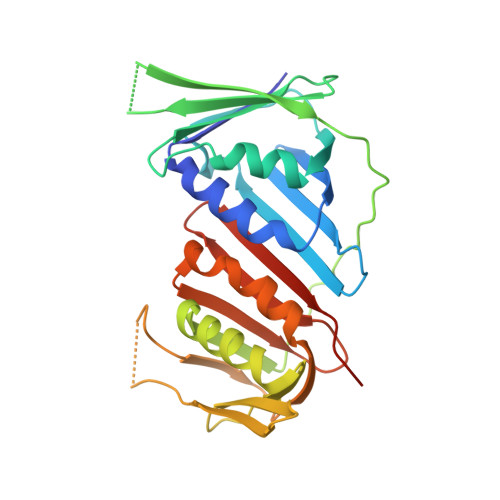
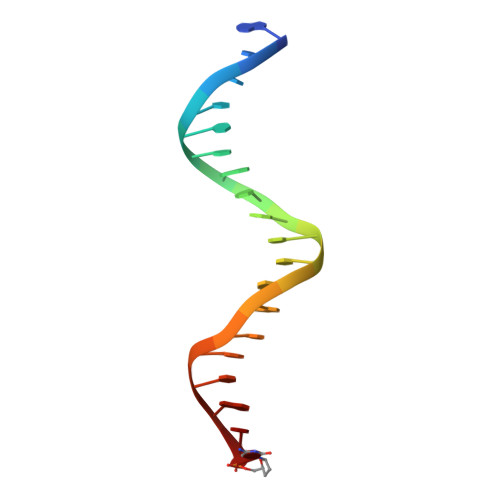
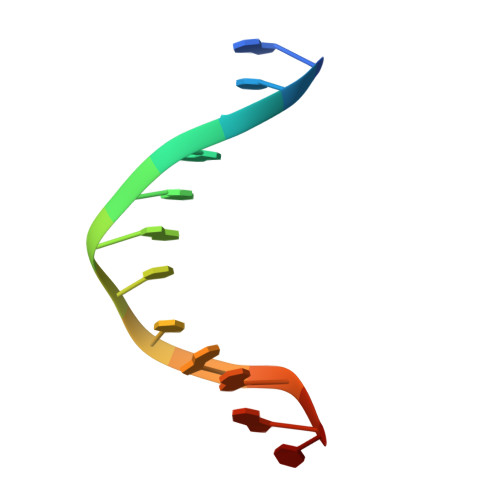
Mechanism of human Lig1 regulation by PCNA in Okazaki fragment sealing.
Blair, K., Tehseen, M., Raducanu, V.S., Shahid, T., Lancey, C., Rashid, F., Creuhet, R., Hamdan, S.M., De Biasio, A.(2022) Nat Commun 13: 7833-7833
- PubMed: 36539424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-35475-z
- Primary Citation Related Structures:
7QNZ, 7QO1, 8B8T - PubMed Abstract:
During lagging strand synthesis, DNA Ligase 1 (Lig1) cooperates with the sliding clamp PCNA to seal the nicks between Okazaki fragments generated by Pol δ and Flap endonuclease 1 (FEN1). We present several cryo-EM structures combined with functional assays, showing that human Lig1 recruits PCNA to nicked DNA using two PCNA-interacting motifs (PIPs) located at its disordered N-terminus (PIP N-term ) and DNA binding domain (PIP DBD ). Once Lig1 and PCNA assemble as two-stack rings encircling DNA, PIP N-term is released from PCNA and only PIP DBD is required for ligation to facilitate the substrate handoff from FEN1. Consistently, we observed that PCNA forms a defined complex with FEN1 and nicked DNA, and it recruits Lig1 to an unoccupied monomer creating a toolbelt that drives the transfer of DNA to Lig1. Collectively, our results provide a structural model on how PCNA regulates FEN1 and Lig1 during Okazaki fragments maturation.
- Leicester Institute of Structural & Chemical Biology and Department of Molecular & Cell Biology, University of Leicester, Lancaster Rd, Leicester, LE1 7HB, UK.
Organizational Affiliation: