Crystal structure of FLT3 T343I in complex with the canonical ligand FL
Pannecoucke, E., Savvides, S.N.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fms-related tyrosine kinase 3 ligand | 155 | Homo sapiens | Mutation(s): 0 Gene Names: FLT3LG | 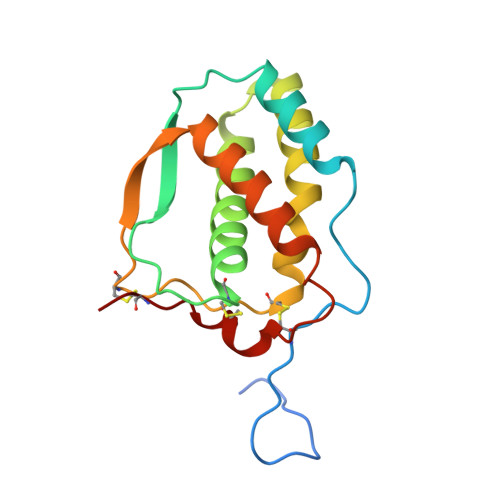 | |
UniProt & NIH Common Fund Data Resources | |||||
GTEx: ENSG00000090554 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P49771 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Receptor-type tyrosine-protein kinase FLT3 | 582 | Homo sapiens | Mutation(s): 2 Gene Names: FLT3, CD135, FLK2, STK1 EC: 2.7.10.1 | 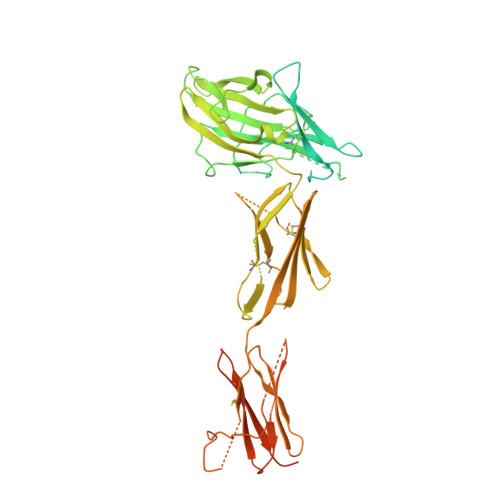 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P36888 GTEx: ENSG00000122025 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P36888 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | Go to GlyGen: P36888-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | I | 3 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G15407YE GlyCosmos: G15407YE GlyGen: G15407YE | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | K [auth E], L [auth F], M [auth F], N [auth H] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 102.428 | α = 105.37 |
| b = 113.377 | β = 109.47 |
| c = 123.225 | γ = 108.22 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| MxCuBE | data collection |
| XDS | data reduction |
| XDS | data scaling |
| Coot | model building |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Research Foundation - Flanders (FWO) | Belgium | -- |