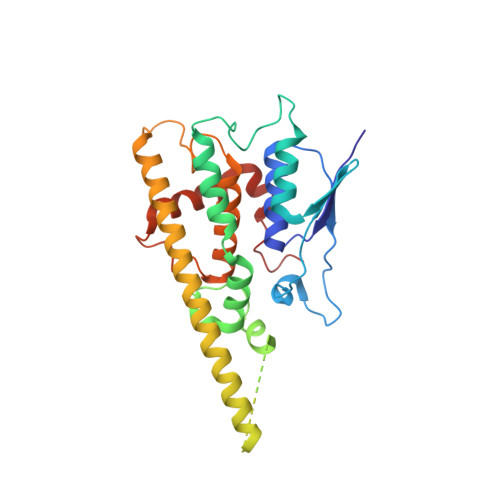
Structural insights into Charcot-Marie-Tooth disease-linked mutations in human GDAP1.
Sutinen, A., Nguyen, G.T.T., Raasakka, A., Muruganandam, G., Loris, R., Ylikallio, E., Tyynismaa, H., Bartesaghi, L., Ruskamo, S., Kursula, P.(2022) FEBS Open Bio 12: 1306-1324
- PubMed: 35509130 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/2211-5463.13422
- Primary Citation Related Structures:
7Q6J, 7Q6K, 7YWD - PubMed Abstract:
Charcot-Marie-Tooth disease (CMT) is the most common inherited peripheral polyneuropathy in humans, and its different subtypes are linked to mutations in dozens of different genes. Mutations in ganglioside-induced differentiation-associated protein 1 (GDAP1) cause two types of CMT, demyelinating CMT4A and axonal CMT2K. The GDAP1-linked CMT genotypes are mainly missense point mutations. Despite clinical profiling and in vivo studies on the mutations, the etiology of GDAP1-linked CMT is poorly understood. Here, we describe the biochemical and structural properties of the Finnish founding CMT2K mutation H123R and CMT2K-linked R120W, both of which are autosomal dominant mutations. The disease variant proteins retain close to normal structure and solution behavior, but both present a significant decrease in thermal stability. Using GDAP1 variant crystal structures, we identify a side-chain interaction network between helices ⍺3, ⍺6, and ⍺7, which is affected by CMT mutations, as well as a hinge in the long helix ⍺6, which is linked to structural flexibility. Structural analysis of GDAP1 indicates that CMT may arise from disruption of specific intra- and intermolecular interaction networks, leading to alterations in GDAP1 structure and stability, and, eventually, insufficient motor and sensory neuron function.
- Faculty of Biochemistry and Molecular Medicine & Biocenter Oulu, University of Oulu, Finland.
Organizational Affiliation: