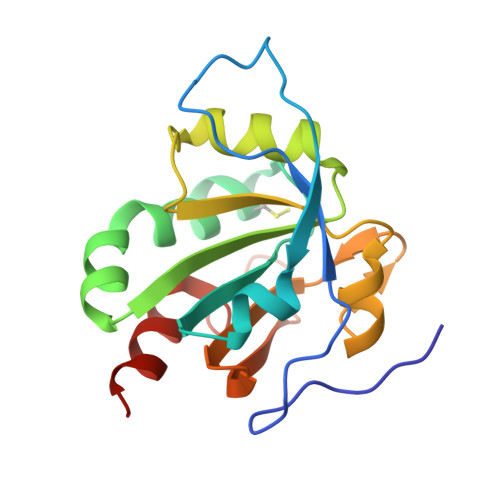
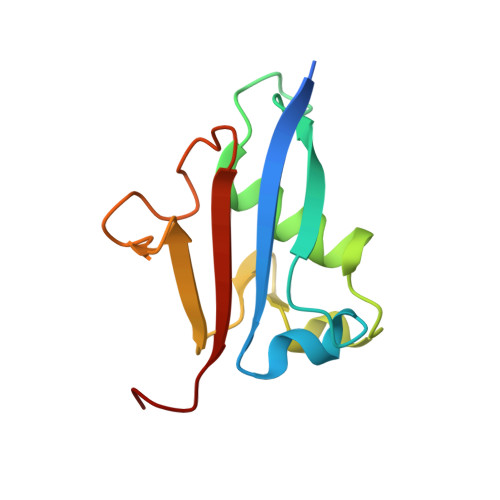
E2/E3-independent ubiquitin-like protein conjugation by Urm1 is directly coupled to cysteine persulfidation.
Ravichandran, K.E., Kaduhr, L., Skupien-Rabian, B., Shvetsova, E., Sokolowski, M., Krutyholowa, R.C., Kwasna, D., Brachmann, C., Lin, S., Guzman Perez, S., Wilk, P., Kosters, M., Grudnik, P., Jankowska, U., Leidel, S.A., Schaffrath, R., Glatt, S.(2022) EMBO J 41: e111318-e111318
- PubMed: 36102610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2022111318
- Primary Citation Related Structures:
7Q5N, 7Q68, 7Q69, 7Q6A - PubMed Abstract:
Post-translational modifications by ubiquitin-like proteins (UBLs) are essential for nearly all cellular processes. Ubiquitin-related modifier 1 (Urm1) is a unique UBL, which plays a key role in tRNA anticodon thiolation as a sulfur carrier protein (SCP) and is linked to the noncanonical E1 enzyme Uba4 (ubiquitin-like protein activator 4). While Urm1 has also been observed to conjugate to target proteins like other UBLs, the molecular mechanism of its attachment remains unknown. Here, we reconstitute the covalent attachment of thiocarboxylated Urm1 to various cellular target proteins in vitro, revealing that, unlike other known UBLs, this process is E2/E3-independent and requires oxidative stress. Furthermore, we present the crystal structures of the peroxiredoxin Ahp1 before and after the covalent attachment of Urm1. Surprisingly, we show that urmylation is accompanied by the transfer of sulfur to cysteine residues in the target proteins, also known as cysteine persulfidation. Our results illustrate the role of the Uba4-Urm1 system as a key evolutionary link between prokaryotic SCPs and the UBL modifications observed in modern eukaryotes.
- Malopolska Centre of Biotechnology (MCB), Jagiellonian University, Krakow, Poland.
Organizational Affiliation: