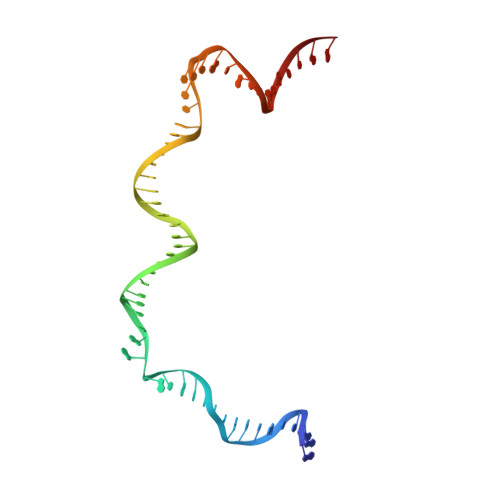
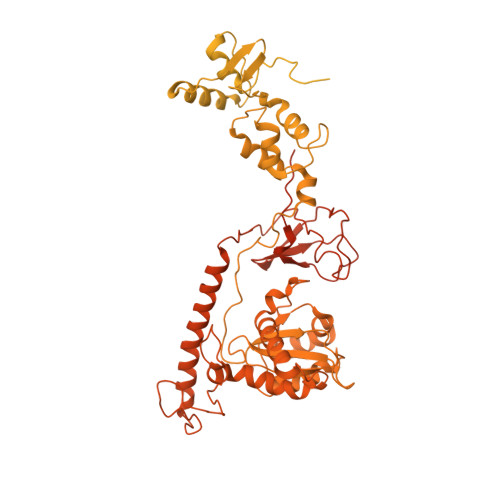
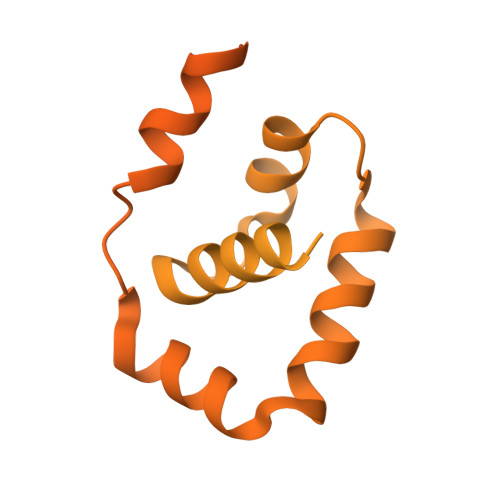
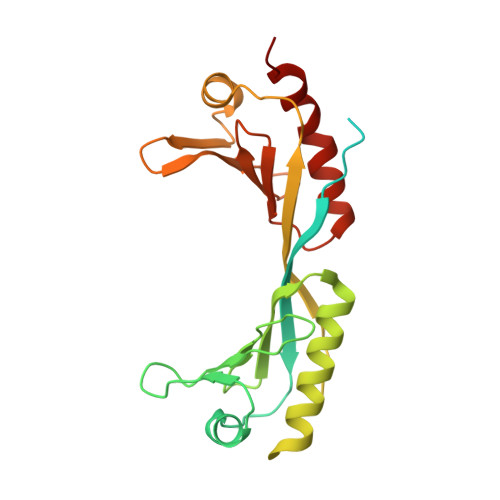
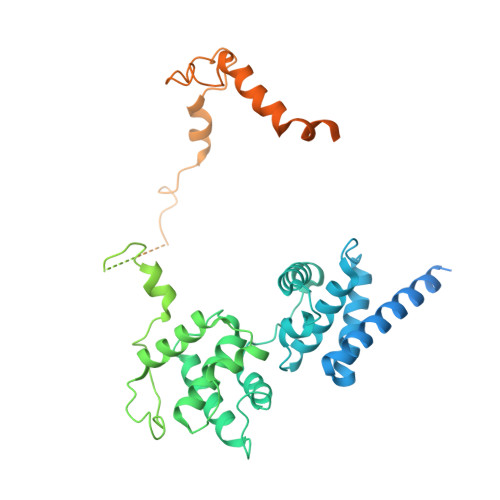
Structural basis of Ty3 retrotransposon integration at RNA Polymerase III-transcribed genes.
Abascal-Palacios, G., Jochem, L., Pla-Prats, C., Beuron, F., Vannini, A.(2021) Nat Commun 12: 6992-6992
- PubMed: 34848735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27338-w
- Primary Citation Related Structures:
7Q5B - PubMed Abstract:
Retrotransposons are endogenous elements that have the ability to mobilise their DNA between different locations in the host genome. The Ty3 retrotransposon integrates with an exquisite specificity in a narrow window upstream of RNA Polymerase (Pol) III-transcribed genes, representing a paradigm for harmless targeted integration. Here we present the cryo-EM reconstruction at 4.0 Å of an active Ty3 strand transfer complex bound to TFIIIB transcription factor and a tRNA gene. The structure unravels the molecular mechanisms underlying Ty3 targeting specificity at Pol III-transcribed genes and sheds light into the architecture of retrotransposon machinery during integration. Ty3 intasome contacts a region of TBP, a subunit of TFIIIB, which is blocked by NC2 transcription regulator in RNA Pol II-transcribed genes. A newly-identified chromodomain on Ty3 integrase interacts with TFIIIB and the tRNA gene, defining with extreme precision the integration site position.
- Division of Structural Biology, The Institute of Cancer Research, London, SW7 3RP, UK. guiller.abascalpalacios@icr.ac.uk.
Organizational Affiliation: