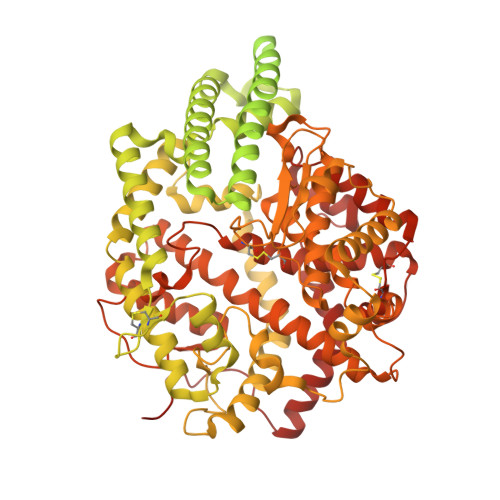
Cryo-EM reveals mechanisms of angiotensin I-converting enzyme allostery and dimerization.
Lubbe, L., Sewell, B.T., Woodward, J.D., Sturrock, E.D.(2022) EMBO J 41: e110550-e110550
- PubMed: 35818993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2021110550
- Primary Citation Related Structures:
7Q3Y, 7Q49, 7Q4C, 7Q4D, 7Q4E - PubMed Abstract:
Hypertension (high blood pressure) is a major risk factor for cardiovascular disease, which is the leading cause of death worldwide. The somatic isoform of angiotensin I-converting enzyme (sACE) plays a critical role in blood pressure regulation, and ACE inhibitors are thus widely used to treat hypertension and cardiovascular disease. Our current understanding of sACE structure, dynamics, function, and inhibition has been limited because truncated, minimally glycosylated forms of sACE are typically used for X-ray crystallography and molecular dynamics simulations. Here, we report the first cryo-EM structures of full-length, glycosylated, soluble sACE (sACE S1211 ). Both monomeric and dimeric forms of the highly flexible apo enzyme were reconstructed from a single dataset. The N- and C-terminal domains of monomeric sACE S1211 were resolved at 3.7 and 4.1 Å, respectively, while the interacting N-terminal domains responsible for dimer formation were resolved at 3.8 Å. Mechanisms are proposed for intradomain hinging, cooperativity, and homodimerization. Furthermore, the observation that both domains were in the open conformation has implications for the design of sACE modulators.
- Department of Integrative Biomedical Sciences, Institute of Infectious Disease and Molecular Medicine, University of Cape Town, Cape Town, South Africa.
Organizational Affiliation: