Methyl 2-Halo-4-Substituted-5-Sulfamoyl-Benzoates as High Affinity and Selective Inhibitors of Carbonic Anhydrase IX.
Zaksauskas, A., Capkauskaite, E., Paketuryte-Latve, V., Smirnov, A., Leitans, J., Kazaks, A., Dvinskis, E., Stancaitis, L., Mickeviciute, A., Jachno, J., Jezepcikas, L., Linkuviene, V., Sakalauskas, A., Manakova, E., Grazulis, S., Matuliene, J., Tars, K., Matulis, D.(2021) Int J Mol Sci 23
- PubMed: 35008553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms23010130
- Primary Citation Related Structures:
7POM, 7PP9, 7PUU, 7PUV, 7PUW, 7Q0C, 7Q0D, 7Q0E - PubMed Abstract:
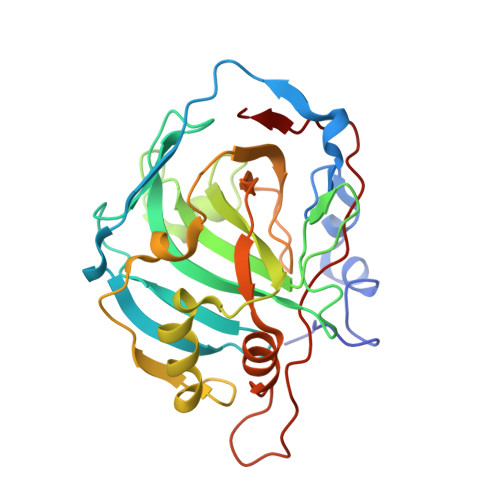
Among the twelve catalytically active carbonic anhydrase isozymes present in the human body, the CAIX is highly overexpressed in various solid tumors. The enzyme acidifies the tumor microenvironment enabling invasion and metastatic processes. Therefore, many attempts have been made to design chemical compounds that would exhibit high affinity and selective binding to CAIX over the remaining eleven catalytically active CA isozymes to limit undesired side effects. It has been postulated that such drugs may have anticancer properties and could be used in tumor treatment. Here we have designed a series of compounds, methyl 5-sulfamoyl-benzoates, which bear a primary sulfonamide group, a well-known marker of CA inhibitors, and determined their affinities for all twelve CA isozymes. Variations of substituents on the benzenesulfonamide ring led to compound 4b , which exhibited an extremely high observed binding affinity to CAIX; the K d was 0.12 nM. The intrinsic dissociation constant, where the binding-linked protonation reactions have been subtracted, reached 0.08 pM. The compound also exhibited more than 100-fold selectivity over the remaining CA isozymes. The X-ray crystallographic structure of compound 3b bound to CAIX showed the structural position, while several structures of compounds bound to other CA isozymes showed structural reasons for compound selectivity towards CAIX. Since this series of compounds possess physicochemical properties suitable for drugs, they may be developed for anticancer therapeutic purposes.
- Department of Biothermodynamics and Drug Design, Institute of Biotechnology, Life Sciences Center, Vilnius University, Saulėtekio al. 7, LT-10257 Vilnius, Lithuania.
Organizational Affiliation: