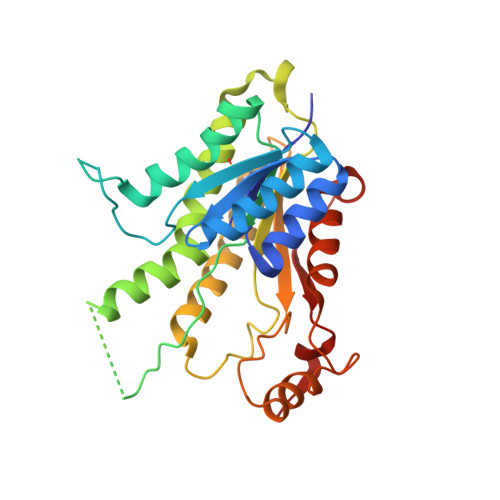
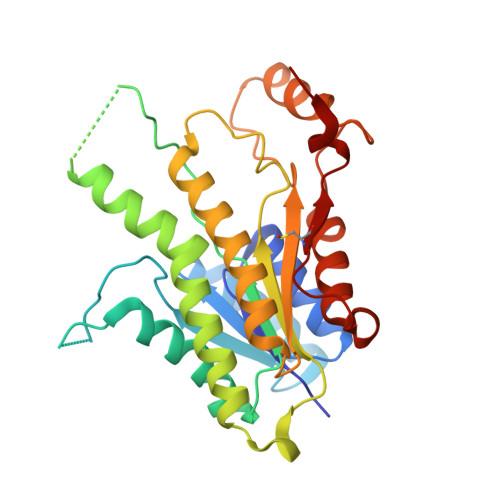
Crystal structure of the ternary complex of Leishmania major pteridine reductase 1 with the cofactor NADP + /NADPH and the substrate folic acid.
Dello Iacono, L., Di Pisa, F., Mangani, S.(2022) Acta Crystallogr F Struct Biol Commun 78: 170-176
- PubMed: 35400669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22002795
- Primary Citation Related Structures:
7PXX - PubMed Abstract:
Pteridine reductase 1 (PTR1) is a key enzyme of the folate pathway in protozoan parasites of the genera Leishmania and Trypanosoma and is a valuable drug target for tropical diseases. This enzyme is able to catalyze the NADPH-dependent reduction of both conjugated (folate) and unconjugated (biopterin) pterins to their tetrahydro forms, starting from oxidized- or dihydro-state substrates. The currently available X-ray structures of Leishmania major PTR1 (LmPTR1) show the enzyme in its unbound, unconjugated substrate-bound (with biopterin derivatives) and inhibitor-bound forms. However, no structure has yet been determined of LmPTR1 bound to a conjugated substrate. Here, the high-resolution crystal structure of LmPTR1 in complex with folic acid is presented and the intermolecular forces that drive the binding of the substrate in the catalytic pocket are described. By expanding the collection of LmPTR1 structures in complex with process intermediates, additional insights into the active-site rearrangements that occur during the catalytic process are provided. In contrast to previous structures with biopterin derivatives, a small but significant difference in the orientation of Asp181 and Tyr194 of the catalytic triad is found. This feature is shared by PTR1 from T. brucei (TbPTR1) in complex with the same substrate molecule and may be informative in deciphering the importance of such residues at the beginning of the catalytic process.
- Department of Biotechnology, Chemistry and Pharmacy, University of Siena, Via Aldo Moro 2, 53100 Siena, Italy.
Organizational Affiliation: