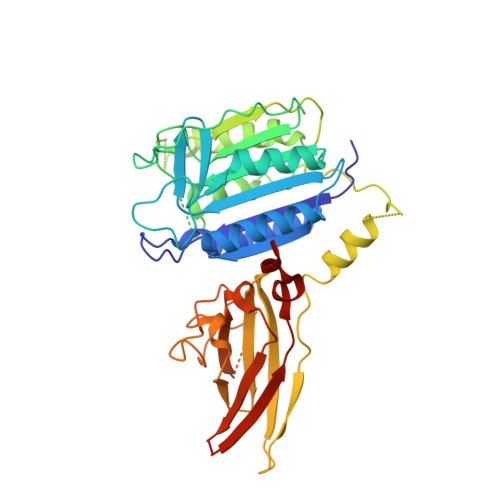
Discovery and optimization of cyclohexane-1,4-diamines as allosteric MALT1 inhibitors.
Schiesser, S., Hajek, P., Pople, H.E., Kack, H., Oster, L., Cox, R.J.(2021) Eur J Med Chem 227: 113925-113925
- PubMed: 34742013 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2021.113925
- Primary Citation Related Structures:
7PAV, 7PAW - PubMed Abstract:
Inhibition of mucosa-associated lymphoid tissue lymphoma translocation protein-1 (MALT1) is a promising strategy to modulate NF-κB signaling, with the potential to treat B-cell lymphoma and autoimmune diseases. We describe the discovery and optimization of (1s,4s)-N,N'-diaryl cyclohexane-1,4-diamines, a novel series of allosteric MALT1 inhibitors, resulting in compound 8 with single digit micromolar cell potency. X-ray analysis confirms that this compound binds to an induced allosteric site in MALT1. Compound 8 is highly selective and has an excellent in vivo rat PK profile with low clearance and high oral bioavailability, making it a promising lead for further optimization.
- Department of Medicinal Chemistry, Research and Early Development, Respiratory & Immunology (R&I), BioPharmaceuticals R&D, AstraZeneca, Pepparedsleden 1, 43183, Mölndal, Sweden. Electronic address: stefan.schiesser@astrazeneca.com.
Organizational Affiliation: