Structural characterization of human neutralizing antibodies against JC and BK polyomavirus
Harprecht, C., Stroeh, L.J., Senn, L., O'Hara, B.A., Nagel, F., Freytag, J., Stiehrof, Y.D., Atwood, W.J., Combaluzier, B., Stehle, T.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Major capsid protein VP1 | 272 | JC polyomavirus | Mutation(s): 0 | 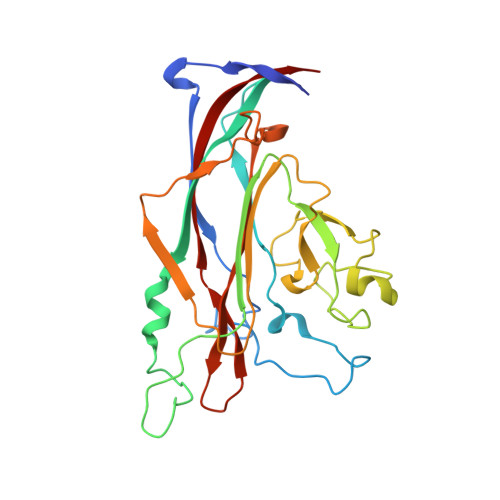 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P03089 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab 98D3 VH | 404 | Homo sapiens | Mutation(s): 0 | 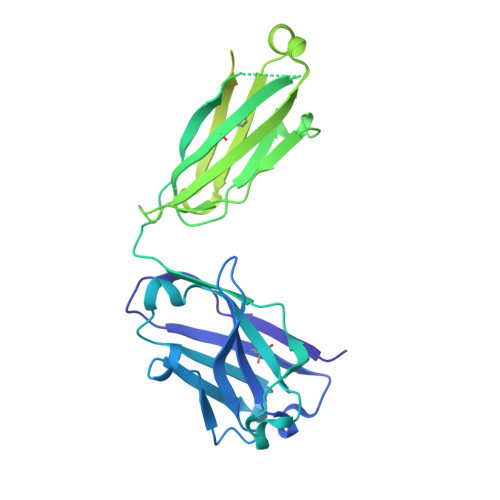 | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab 98D3 VL | 212 | Homo sapiens | Mutation(s): 0 | 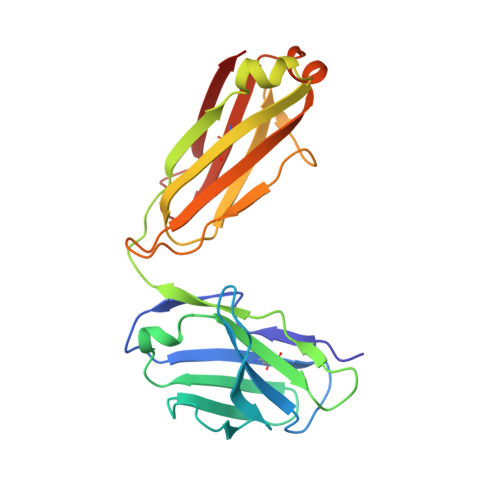 | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 170.97 | α = 90 |
| b = 202.31 | β = 90 |
| c = 251.66 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| German Research Foundation (DFG) | Germany | FOR2327 |