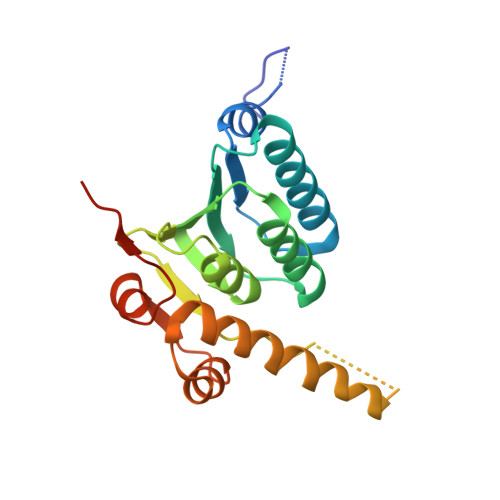
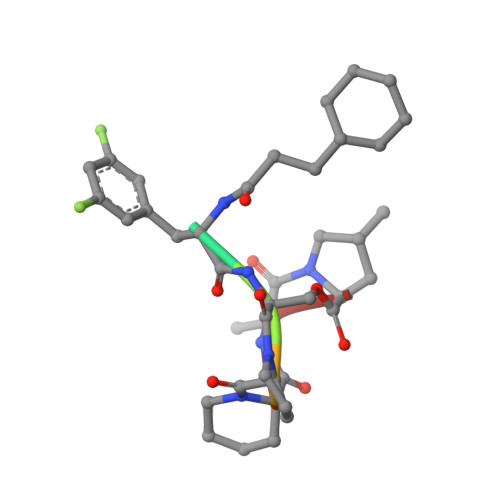
Structural insights into ClpP protease side exit pore-opening by a pH drop coupled with substrate hydrolysis.
Kim, L., Lee, B.G., Kim, M., Kim, M.K., Kwon, D.H., Kim, H., Brotz-Oesterhelt, H., Roh, S.H., Song, H.K.(2022) EMBO J 41: e109755-e109755
- PubMed: 35593068 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2021109755
- Primary Citation Related Structures:
7FEP, 7FEQ, 7FER, 7FES, 7P80, 7P81 - PubMed Abstract:
The ClpP serine peptidase is a tetradecameric degradation molecular machine involved in many physiological processes. It becomes a competent ATP-dependent protease when coupled with Clp-ATPases. Small chemical compounds, acyldepsipeptides (ADEPs), are known to cause the dysregulation and activation of ClpP without ATPases and have potential as novel antibiotics. Previously, structural studies of ClpP from various species revealed its structural details, conformational changes, and activation mechanism. Although product release through side exit pores has been proposed, the detailed driving force for product release remains elusive. Herein, we report crystal structures of ClpP from Bacillus subtilis (BsClpP) in unforeseen ADEP-bound states. Cryo-electron microscopy structures of BsClpP revealed various conformational states under different pH conditions. To understand the conformational change required for product release, we investigated the relationship between substrate hydrolysis and the pH-lowering process. The production of hydrolyzed peptides from acidic and basic substrates by proteinase K and BsClpP lowered the pH values. Our data, together with those of previous findings, provide insight into the molecular mechanism of product release by the ClpP self-compartmentalizing protease.
- Department of Life Sciences, Korea University, Seoul, South Korea.
Organizational Affiliation: