Medicinal Au(I) compounds targeting urease as prospective antimicrobial agents: unveiling the structural basis for enzyme inhibition.
Mazzei, L., Massai, L., Cianci, M., Messori, L., Ciurli, S.(2021) Dalton Trans 50: 14444-14452
- PubMed: 34585201 Search on PubMed
- DOI: https://doi.org/10.1039/d1dt02488d
- Primary Citation Related Structures:
7P7N, 7P7O - PubMed Abstract:
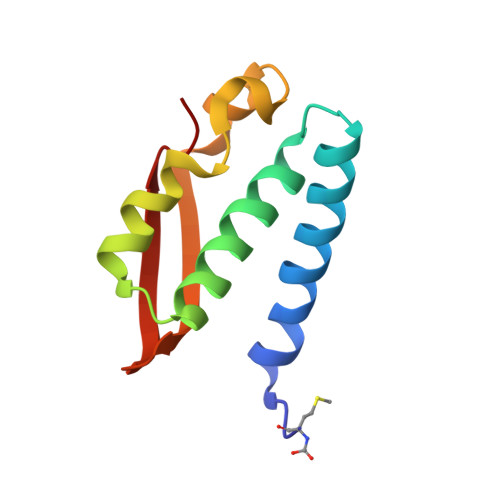
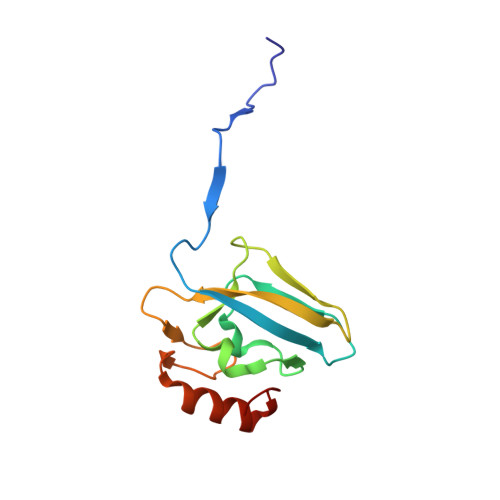
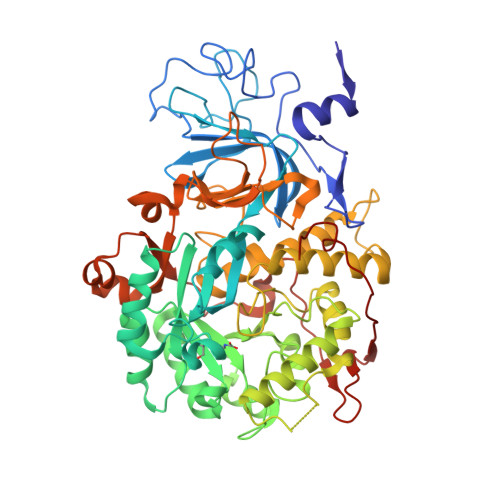
A few gold compounds were recently found to show antimicrobial properties in vitro , holding great promise for the discovery of new drugs to overcome antibiotic resistance. Here, the inhibition of the bacterial virulence factor urease by four Au(I)-compounds, namely Au(PEt 3 )Cl, Au(PEt 3 )Br, Au(PEt 3 )I and [Au(PEt 3 ) 2 ]Cl, obtained from the antiarthritic Au(I)-drug Auranofin and earlier reported to act as antimicrobials, is investigated. The three monophosphino Au(I) complexes showed IC 50 values in the 30-100 nM range, while the diphosphino Au(I) complex, though being less active, still showed a IC 50 value of 7 μM. The structural basis for this inhibition was provided by solving the crystal structures of urease co-crystallized with Au(PEt 3 )I and [Au(PEt 3 ) 2 ]Cl: at least two Au(I) ions bind the enzyme in a flap domain involved in the catalysis, thus obliterating enzyme activity. Peculiar changes observed in the two structures reveal implications for the mechanism of soft metal binding and enzyme inactivation.
- Laboratory of Bioinorganic Chemistry, Department of Pharmacy and Biotechnology (FaBiT), University of Bologna, Via Giuseppe Fanin 40, I-40127 Bologna, Italy. luca.mazzei2@unibo.it.
Organizational Affiliation: