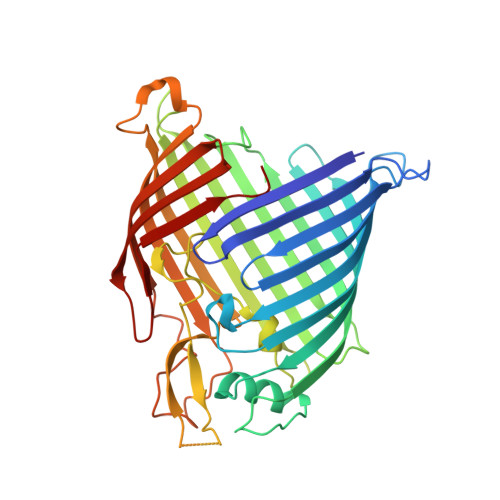
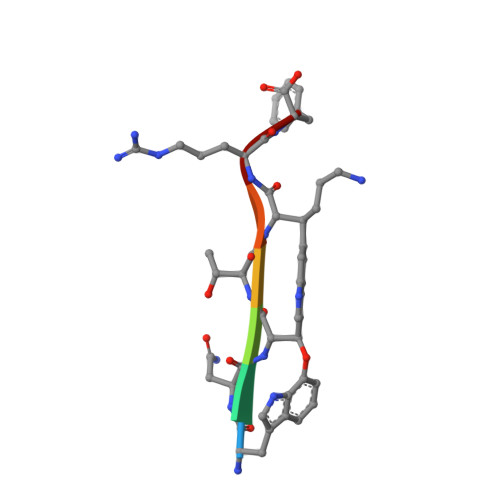
Mutasynthetic Production and Antimicrobial Characterization of Darobactin Analogs.
Bohringer, N., Green, R., Liu, Y., Mettal, U., Marner, M., Modaresi, S.M., Jakob, R.P., Wuisan, Z.G., Maier, T., Iinishi, A., Hiller, S., Lewis, K., Schaberle, T.F.(2021) Microbiol Spectr 9: e0153521-e0153521
- PubMed: 34937193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/spectrum.01535-21
- Primary Citation Related Structures:
7P1C - PubMed Abstract:
There is great need for therapeutics against multidrug-resistant, Gram-negative bacterial pathogens. Recently, darobactin A, a novel bicyclic heptapeptide that selectively kills Gram-negative bacteria by targeting the outer membrane protein BamA, was discovered. Its efficacy was proven in animal infection models of Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa, thus promoting darobactin A as a promising lead compound. Originally discovered from members of the nematode-symbiotic genus Photorhabdus , the biosynthetic gene cluster (BGC) encoding the synthesis of darobactin A can also be found in other members of the class Gammaproteobacteria . Therein, the precursor peptides DarB to -F, which differ in their core sequence from darobactin A, were identified in silico . Even though production of these analogs was not observed in the putative producer strains, we were able to generate them by mutasynthetic derivatization of a heterologous expression system. The analogs generated were isolated and tested for their bioactivity. The most potent compound, darobactin B, was used for cocrystallization with the target BamA, revealing a binding site identical to that of darobactin A. Despite its potency, darobactin B did not exhibit cytotoxicity, and it was slightly more active against Acinetobacter baumannii isolates than darobactin A. Furthermore, we evaluated the plasma protein binding of darobactin A and B, indicating their different pharmacokinetic properties. This is the first report on new members of this new antibiotic class, which is likely to expand to several promising therapeutic candidates. IMPORTANCE Therapeutic options to combat Gram-negative bacterial pathogens are dwindling with increasing antibiotic resistance. This study presents a proof of concept for the heterologous-expression approach to expand on the novel antibiotic class of darobactins and to generate analogs with different activities and pharmacokinetic properties. In combination with the structural data of the target BamA, this approach may contribute to structure-activity relationship (SAR) data to optimize inhibitors of this essential outer membrane protein of Gram-negative pathogens.
- Justus-Liebig-University Gießen, Gießen, Germany.
Organizational Affiliation: