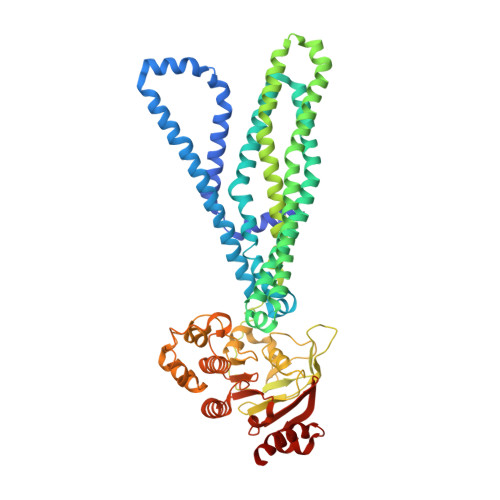
Substrate-bound and substrate-free outward-facing structures of a multidrug ABC exporter.
Chaptal, V., Zampieri, V., Wiseman, B., Orelle, C., Martin, J., Nguyen, K.A., Gobet, A., Di Cesare, M., Magnard, S., Javed, W., Eid, J., Kilburg, A., Peuchmaur, M., Marcoux, J., Monticelli, L., Hogbom, M., Schoehn, G., Jault, J.M., Boumendjel, A., Falson, P.(2022) Sci Adv 8: eabg9215-eabg9215
- PubMed: 35080979 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abg9215
- Primary Citation Related Structures:
6R72, 6R81, 7BG4, 7OW8 - PubMed Abstract:
Multidrug ABC transporters translocate drugs across membranes by a mechanism for which the molecular features of drug release are so far unknown. Here, we resolved three ATP-Mg 2+ -bound outward-facing conformations of the Bacillus subtilis (homodimeric) BmrA by x-ray crystallography and single-particle cryo-electron microscopy (EM) in detergent solution, one of them with rhodamine 6G (R6G), a substrate exported by BmrA when overexpressed in B. subtilis . Two R6G molecules bind to the drug-binding cavity at the level of the outer leaflet, between transmembrane (TM) helices 1-2 of one monomer and TM5'-6' of the other. They induce a rearrangement of TM1-2, highlighting a local flexibility that we confirmed by hydrogen/deuterium exchange and molecular dynamics simulations. In the absence of R6G, simulations show a fast postrelease occlusion of the cavity driven by hydrophobicity, while when present, R6G can move within the cavity, maintaining it open.
- Drug Resistance and Membrane Proteins Group, Molecular Microbiology and Structural Biochemistry Laboratory, CNRS UMR 5086, University of Lyon, IBCP, 7, passage du Vercors, 69367 Lyon, France.
Organizational Affiliation: