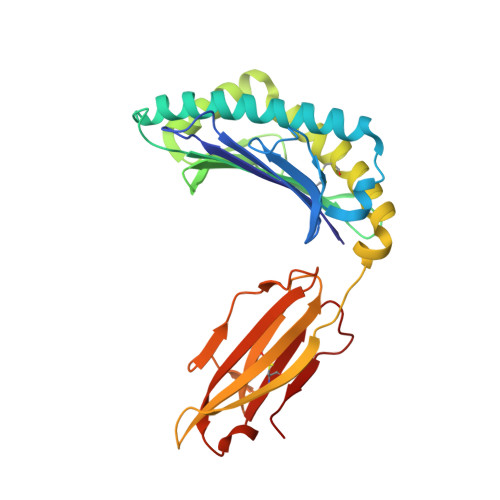
Therapeutic high affinity T cell receptor targeting a KRASG12D cancer neoantigen
Poole, A., Karuppiah, V., Hartt, A., Haidar, J.N., Moureau, S., Dobrzycki, T., Hayes, C., Rowley, C., Dias, J., Harper, S., Barnbrook, K., Hock, M., Coles, C., Yang, W., Aleksic, M., Lin, A.B., Robinson, R., Dukes, J.D., Liddy, N., Van der Kamp, M., Plowman, G.D., Vuidepot, A., Cole, D.K., Whale, A.D., Chillakuri, C.(2022) Nat Commun 13: 5333