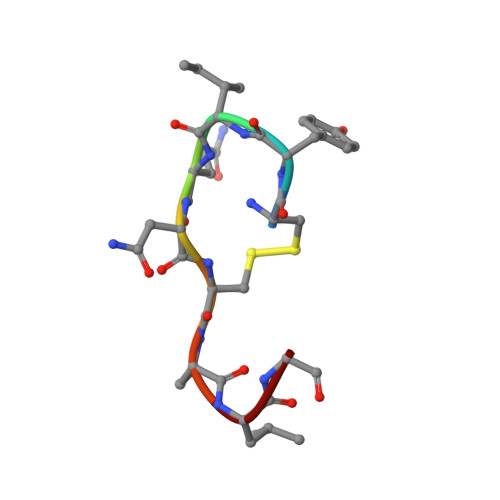
Determining the structure and binding mechanism of oxytocin-Cu 2+ complex using paramagnetic relaxation enhancement NMR analysis.
Alshanski, I., Shalev, D.E., Yitzchaik, S., Hurevich, M.(2021) J Biol Inorg Chem 26: 809-815
- PubMed: 34459989 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00775-021-01897-1
- Primary Citation Related Structures:
7OFG, 7OTD - PubMed Abstract:
Oxytocin is a neuropeptide that binds copper ions in nature. The structure of oxytocin in interaction with Cu 2+ was determined here by NMR, showing which atoms of the peptide are involved in binding. Paramagnetic relaxation enhancement NMR analyses indicated a binding mechanism where the amino terminus was required for binding and subsequently Tyr2, Ile3 and Gln4 bound in that order. The aromatic ring of Tyr2 formed a π-cation interaction with Cu 2+ . Oxytocin copper complex structure revealed by paramagnetic relaxation enhancement NMR analyses.
- Institute of Chemistry and Center for Nanoscience and Nanotechnology, The Hebrew University of Jerusalem, Edmond J. Safra Campus, Givat Ram, 91904, Jerusalem, Israel.
Organizational Affiliation: