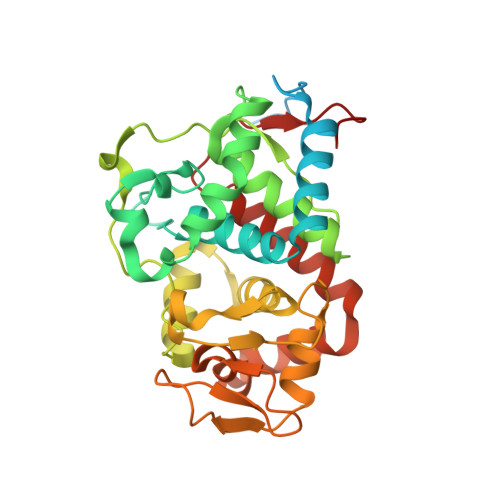
Crystal structure of Trypanosoma cruzi heme peroxidase and characterization of its substrate specificity and compound I intermediate.
Freeman, S.L., Skafar, V., Kwon, H., Fielding, A.J., Moody, P.C.E., Martinez, A., Issoglio, F.M., Inchausti, L., Smircich, P., Zeida, A., Piacenza, L., Radi, R., Raven, E.L.(2022) J Biological Chem 298: 102204-102204
- PubMed: 35772495 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102204
- Primary Citation Related Structures:
7OPT, 7OQR - PubMed Abstract:
The protozoan parasite Trypanosoma cruzi is the causative agent of American trypanosomiasis, otherwise known as Chagas disease. To survive in the host, the T. cruzi parasite needs antioxidant defense systems. One of these is a hybrid heme peroxidase, the T. cruzi ascorbate peroxidase-cytochrome c peroxidase enzyme (TcAPx-CcP). TcAPx-CcP has high sequence identity to members of the class I peroxidase family, notably ascorbate peroxidase (APX) and cytochrome c peroxidase (CcP), as well as a mitochondrial peroxidase from Leishmania major (LmP). The aim of this work was to solve the structure and examine the reactivity of the TcAPx-CcP enzyme. Low temperature electron paramagnetic resonance spectra support the formation of an exchange-coupled [Fe(IV)=O Trp 233 •+ ] compound I radical species, analogous to that used in CcP and LmP. We demonstrate that TcAPx-CcP is similar in overall structure to APX and CcP, but there are differences in the substrate-binding regions. Furthermore, the electron transfer pathway from cytochrome c to the heme in CcP and LmP is preserved in the TcAPx-CcP structure. Integration of steady state kinetic experiments, molecular dynamic simulations, and bioinformatic analyses indicates that TcAPx-CcP preferentially oxidizes cytochrome c but is still competent for oxidization of ascorbate. The results reveal that TcAPx-CcP is a credible cytochrome c peroxidase, which can also bind and use ascorbate in host cells, where concentrations are in the millimolar range. Thus, kinetically and functionally TcAPx-CcP can be considered a hybrid peroxidase.
- School of Chemistry, University of Bristol, Bristol, United Kingdom.
Organizational Affiliation: