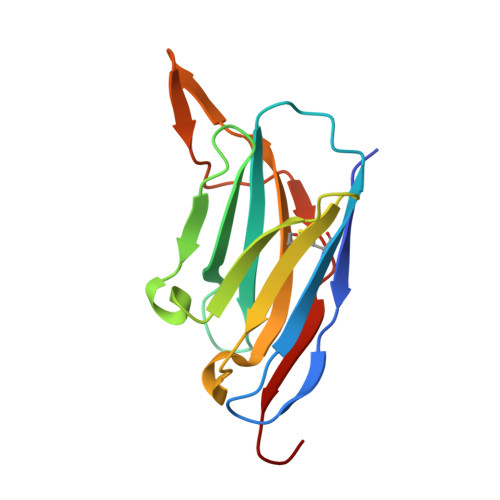
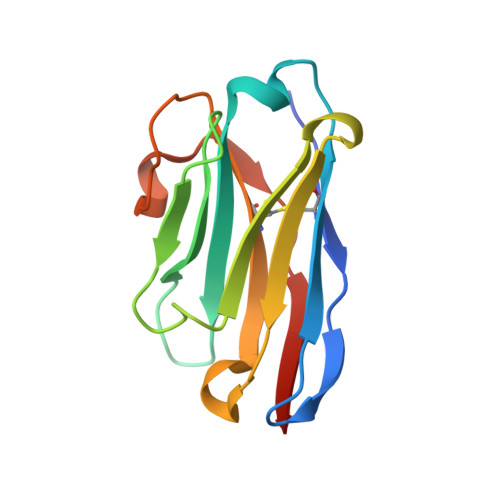
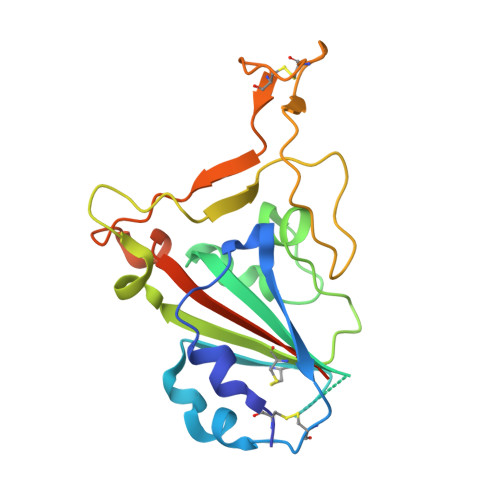
Neutralization of SARS-CoV-2 by highly potent, hyperthermostable, and mutation-tolerant nanobodies.
Guttler, T., Aksu, M., Dickmanns, A., Stegmann, K.M., Gregor, K., Rees, R., Taxer, W., Rymarenko, O., Schunemann, J., Dienemann, C., Gunkel, P., Mussil, B., Krull, J., Teichmann, U., Gross, U., Cordes, V.C., Dobbelstein, M., Gorlich, D.(2021) EMBO J 40: e107985-e107985
- PubMed: 34302370 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2021107985
- Primary Citation Related Structures:
7OLZ, 7ON5 - PubMed Abstract:
Monoclonal anti-SARS-CoV-2 immunoglobulins represent a treatment option for COVID-19. However, their production in mammalian cells is not scalable to meet the global demand. Single-domain (VHH) antibodies (also called nanobodies) provide an alternative suitable for microbial production. Using alpaca immune libraries against the receptor-binding domain (RBD) of the SARS-CoV-2 Spike protein, we isolated 45 infection-blocking VHH antibodies. These include nanobodies that can withstand 95°C. The most effective VHH antibody neutralizes SARS-CoV-2 at 17-50 pM concentration (0.2-0.7 µg per liter), binds the open and closed states of the Spike, and shows a tight RBD interaction in the X-ray and cryo-EM structures. The best VHH trimers neutralize even at 40 ng per liter. We constructed nanobody tandems and identified nanobody monomers that tolerate the K417N/T, E484K, N501Y, and L452R immune-escape mutations found in the Alpha, Beta, Gamma, Epsilon, Iota, and Delta/Kappa lineages. We also demonstrate neutralization of the Beta strain at low-picomolar VHH concentrations. We further discovered VHH antibodies that enforce native folding of the RBD in the E. coli cytosol, where its folding normally fails. Such "fold-promoting" nanobodies may allow for simplified production of vaccines and their adaptation to viral escape-mutations.
- Department of Cellular Logistics, Max Planck Institute for Biophysical Chemistry, Göttingen, Germany.
Organizational Affiliation: