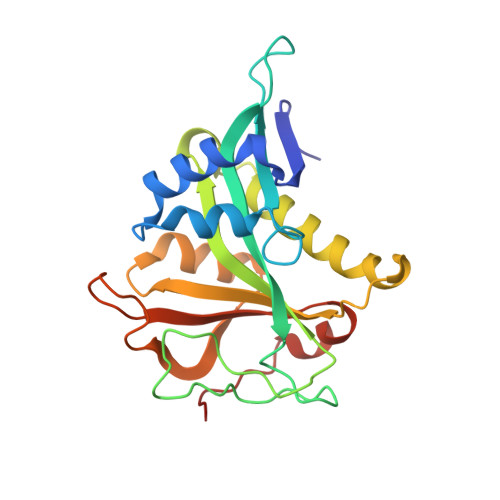
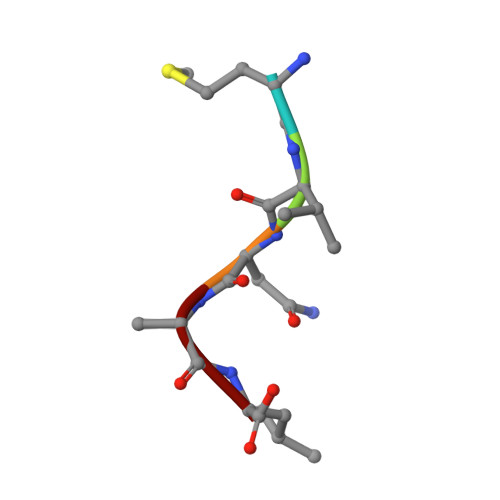
Extended N-Terminal Acetyltransferase Naa50 in Filamentous Fungi Adds to Naa50 Diversity.
Weidenhausen, J., Kopp, J., Ruger-Herreros, C., Stein, F., Haberkant, P., Lapouge, K., Sinning, I.(2022) Int J Mol Sci 23
- PubMed: 36142717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms231810805
- Primary Citation Related Structures:
7OJU - PubMed Abstract:
Most eukaryotic proteins are N-terminally acetylated by a set of Nα acetyltransferases (NATs). This ancient and ubiquitous modification plays a fundamental role in protein homeostasis, while mutations are linked to human diseases and phenotypic defects. In particular, Naa50 features species-specific differences, as it is inactive in yeast but active in higher eukaryotes. Together with NatA, it engages in NatE complex formation for cotranslational acetylation. Here, we report Naa50 homologs from the filamentous fungi Chaetomium thermophilum and Neurospora crassa with significant N- and C-terminal extensions to the conserved GNAT domain. Structural and biochemical analyses show that Ct Naa50 shares the GNAT structure and substrate specificity with other homologs. However, in contrast to previously analyzed Naa50 proteins, it does not form NatE. The elongated N-terminus increases Naa50 thermostability and binds to dynein light chain protein 1, while our data suggest that conserved positive patches in the C-terminus allow for ribosome binding independent of NatA. Our study provides new insights into the many facets of Naa50 and highlights the diversification of NATs during evolution.
- Heidelberg University Biochemistry Center (BZH), 69120 Heidelberg, Germany.
Organizational Affiliation: