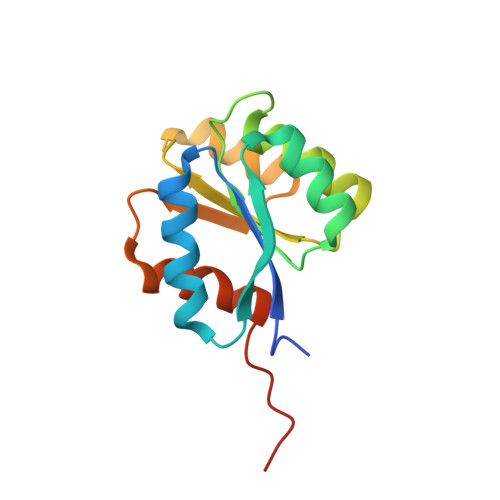
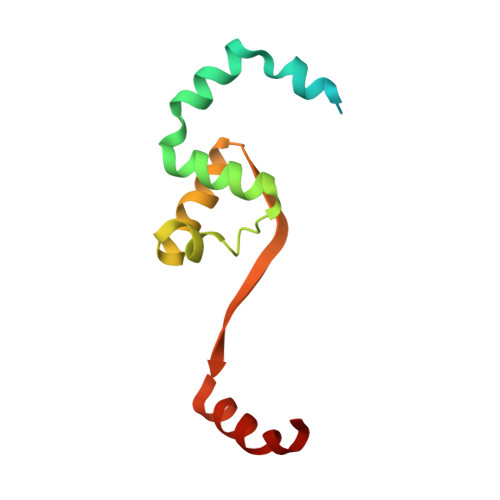
Structural insights into the mechanism of archaellar rotational switching.
Altegoer, F., Quax, T.E.F., Weiland, P., Nussbaum, P., Giammarinaro, P.I., Patro, M., Li, Z., Oesterhelt, D., Grininger, M., Albers, S.V., Bange, G.(2022) Nat Commun 13: 2857-2857
- PubMed: 35606361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30358-9
- Primary Citation Related Structures:
7OD9, 7OVP - PubMed Abstract:
Signal transduction via phosphorylated CheY towards the flagellum and the archaellum involves a conserved mechanism of CheY phosphorylation and subsequent conformational changes within CheY. This mechanism is conserved among bacteria and archaea, despite substantial differences in the composition and architecture of archaellum and flagellum, respectively. Phosphorylated CheY has higher affinity towards the bacterial C-ring and its binding leads to conformational changes in the flagellar motor and subsequent rotational switching of the flagellum. In archaea, the adaptor protein CheF resides at the cytoplasmic face of the archaeal C-ring formed by the proteins ArlCDE and interacts with phosphorylated CheY. While the mechanism of CheY binding to the C-ring is well-studied in bacteria, the role of CheF in archaea remains enigmatic and mechanistic insights are absent. Here, we have determined the atomic structures of CheF alone and in complex with activated CheY by X-ray crystallography. CheF forms an elongated dimer with a twisted architecture. We show that CheY binds to the C-terminal tail domain of CheF leading to slight conformational changes within CheF. Our structural, biochemical and genetic analyses reveal the mechanistic basis for CheY binding to CheF and allow us to propose a model for rotational switching of the archaellum.
- Philipps-University Marburg, Center for Synthetic Microbiology (SYNMIKRO) & Faculty of Chemistry, Karl-von-Frisch-Straße 14, 35043, Marburg, Germany. altegoer@hhu.de.
Organizational Affiliation: