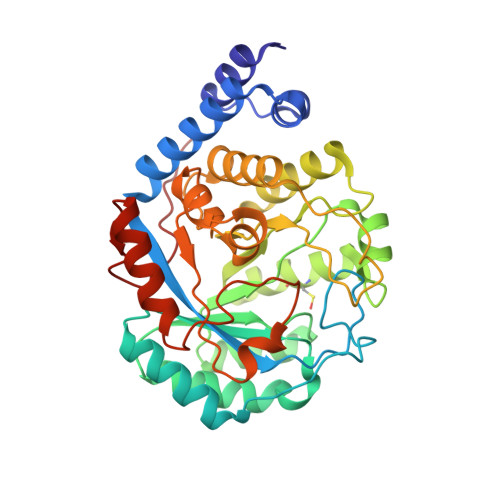
Crystal Structure of the [FeFe]-Hydrogenase Maturase HydE Bound to Complex-B.
Rohac, R., Martin, L., Liu, L., Basu, D., Tao, L., Britt, R.D., Rauchfuss, T.B., Nicolet, Y.(2021) J Am Chem Soc 143: 8499-8508
- PubMed: 34048236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.1c03367
- Primary Citation Related Structures:
7O1O, 7O1P, 7O1S, 7O1T, 7O25, 7O26 - PubMed Abstract:
[FeFe]-hydrogenases use a unique organometallic complex, termed the H cluster, to reversibly convert H 2 into protons and low-potential electrons. It can be best described as a [Fe 4 S 4 ] cluster coupled to a unique [2Fe] H center where the reaction actually takes place. The latter corresponds to two iron atoms, each of which is bound by one CN - ligand and one CO ligand. The two iron atoms are connected by a unique azadithiolate molecule ( - S-CH 2 -NH-CH 2 -S - ) and an additional bridging CO. This [2Fe] H center is built stepwise thanks to the well-orchestrated action of maturating enzymes that belong to the Hyd machinery. Among them, HydG converts l-tyrosine into CO and CN - to produce a unique l-cysteine-Fe(CO) 2 CN species termed complex-B. Very recently, HydE was shown to perform radical-based chemistry using synthetic complex-B as a substrate. Here we report the high-resolution crystal structure that establishes the identity of the complex-B-bound HydE. By triggering the reaction prior to crystallization, we trapped a new five-coordinate Fe species, supporting the proposal that HydE performs complex modifications of complex-B to produce a monomeric "SFe(CO) 2 CN" precursor to the [2Fe] H center. Substrate access, product release, and intermediate transfer are also discussed.
- Univ. Grenoble Alpes, CEA, CNRS, IBS, Metalloproteins Unit, F-38000 Grenoble, France.
Organizational Affiliation: