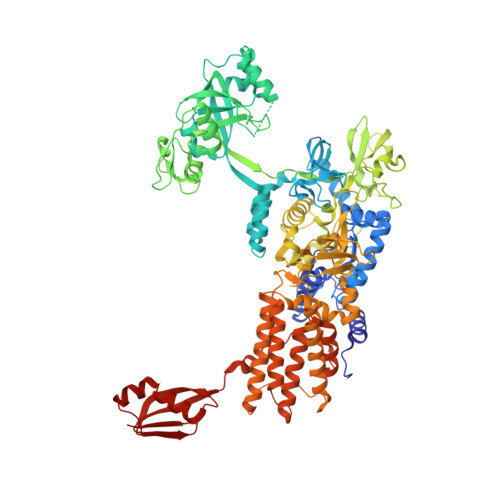
Partitioning of the initial catalytic steps of leucyl-tRNA synthetase is driven by an active site peptide-plane flip.
Pang, L., Zanki, V., Strelkov, S.V., Van Aerschot, A., Gruic-Sovulj, I., Weeks, S.D.(2022) Commun Biol 5: 883-883
- PubMed: 36038645 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-03825-8
- Primary Citation Related Structures:
7NTY, 7NTZ, 7NU0, 7NU1, 7NU2, 7NU3, 7NU4, 7NU5, 7NU6, 7NU7, 7NU8, 7NU9, 7NUA, 7NUB, 7NUC - PubMed Abstract:
To correctly aminoacylate tRNA Leu , leucyl-tRNA synthetase (LeuRS) catalyzes three reactions: activation of leucine by ATP to form leucyl-adenylate (Leu-AMP), transfer of this amino acid to tRNA Leu and post-transfer editing of any mischarged product. Although LeuRS has been well characterized biochemically, detailed structural information is currently only available for the latter two stages of catalysis. We have solved crystal structures for all enzymatic states of Neisseria gonorrhoeae LeuRS during Leu-AMP formation. These show a cycle of dramatic conformational changes, involving multiple domains, and correlate with an energetically unfavorable peptide-plane flip observed in the active site of the pre-transition state structure. Biochemical analyses, combined with mutant structural studies, reveal that this backbone distortion acts as a trigger, temporally compartmentalizing the first two catalytic steps. These results unveil the remarkable effect of this small structural alteration on the global dynamics and activity of the enzyme.
- Biocrystallography, Department of Pharmaceutical and Pharmacological Sciences, KU Leuven, Herestraat 49 - Box 822, 3000, Leuven, Belgium.
Organizational Affiliation: