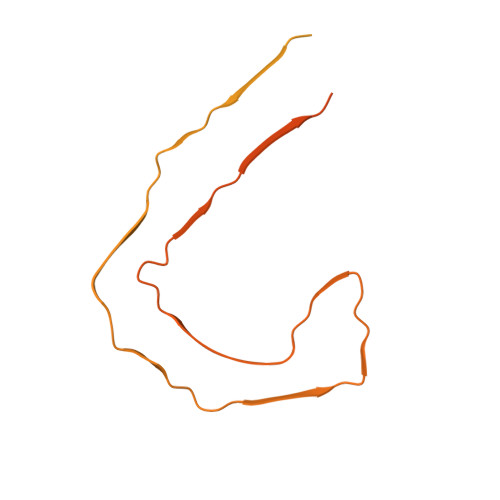
Cryo-EM structures of tau filaments from Alzheimer's disease with PET ligand APN-1607.
Shi, Y., Murzin, A.G., Falcon, B., Epstein, A., Machin, J., Tempest, P., Newell, K.L., Vidal, R., Garringer, H.J., Sahara, N., Higuchi, M., Ghetti, B., Jang, M.K., Scheres, S.H.W., Goedert, M.(2021) Acta Neuropathol 141: 697-708
- PubMed: 33723967 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00401-021-02294-3
- Primary Citation Related Structures:
7NRQ, 7NRS, 7NRT, 7NRV, 7NRX - PubMed Abstract:
Tau and Aβ assemblies of Alzheimer's disease (AD) can be visualized in living subjects using positron emission tomography (PET). Tau assemblies comprise paired helical and straight filaments (PHFs and SFs). APN-1607 (PM-PBB3) is a recently described PET ligand for AD and other tau proteinopathies. Since it is not known where in the tau folds PET ligands bind, we used electron cryo-microscopy (cryo-EM) to determine the binding sites of APN-1607 in the Alzheimer fold. We identified two major sites in the β-helix of PHFs and SFs and a third major site in the C-shaped cavity of SFs. In addition, we report that tau filaments from posterior cortical atrophy (PCA) and primary age-related tauopathy (PART) are identical to those from AD. In support, fluorescence labelling showed binding of APN-1607 to intraneuronal inclusions in AD, PART and PCA. Knowledge of the binding modes of APN-1607 to tau filaments may lead to the development of new ligands with increased specificity and binding activity. We show that cryo-EM can be used to identify the binding sites of small molecules in amyloid filaments.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: