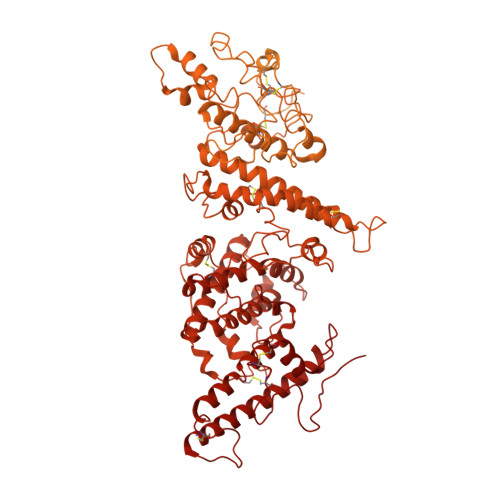
Cryo-EM reveals the architecture of placental malaria VAR2CSA and provides molecular insight into chondroitin sulfate binding.
Wang, K., Dagil, R., Lavstsen, T., Misra, S.K., Spliid, C.B., Wang, Y., Gustavsson, T., Sandoval, D.R., Vidal-Calvo, E.E., Choudhary, S., Agerbaek, M.O., Lindorff-Larsen, K., Nielsen, M.A., Theander, T.G., Sharp, J.S., Clausen, T.M., Gourdon, P., Salanti, A.(2021) Nat Commun 12: 2956-2956
- PubMed: 34011972 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-23254-1
- Primary Citation Related Structures:
7B52, 7B54, 7NNH - PubMed Abstract:
Placental malaria can have severe consequences for both mother and child and effective vaccines are lacking. Parasite-infected red blood cells sequester in the placenta through interaction between parasite-expressed protein VAR2CSA and the glycosaminoglycan chondroitin sulfate A (CS) abundantly present in the intervillous space. Here, we report cryo-EM structures of the VAR2CSA ectodomain at up to 3.1 Å resolution revealing an overall V-shaped architecture and a complex domain organization. Notably, the surface displays a single significantly electropositive patch, compatible with binding of negatively charged CS. Using molecular docking and molecular dynamics simulations as well as comparative hydroxyl radical protein foot-printing of VAR2CSA in complex with placental CS, we identify the CS-binding groove, intersecting with the positively charged patch of the central VAR2CSA structure. We identify distinctive conserved structural features upholding the macro-molecular domain complex and CS binding capacity of VAR2CSA as well as divergent elements possibly allowing immune escape at or near the CS binding site. These observations will support rational design of second-generation placental malaria vaccines.
- Department of Biomedical Sciences, University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: