Functional development of a V3/glycan-specific broadly neutralizing antibody isolated from a case of HIV superinfection.
Shipley, M.M., Mangala Prasad, V., Doepker, L.E., Dingens, A.S., Ralph, D.K., Harkins, E., Dhar, A., Arenz, D., Chohan, V., Weight, H., Mandaliya, K., Bloom, J.D., Matsen Iv, F., Lee, K.K., Overbaugh, J.M.(2021) Elife 10
- PubMed: 34263727 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.68110
- Primary Citation Related Structures:
7N65 - PubMed Abstract:
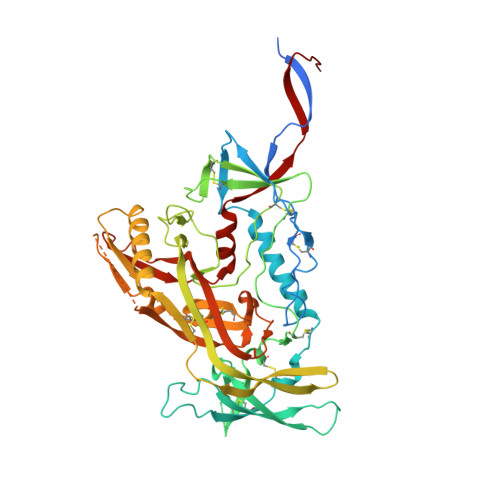
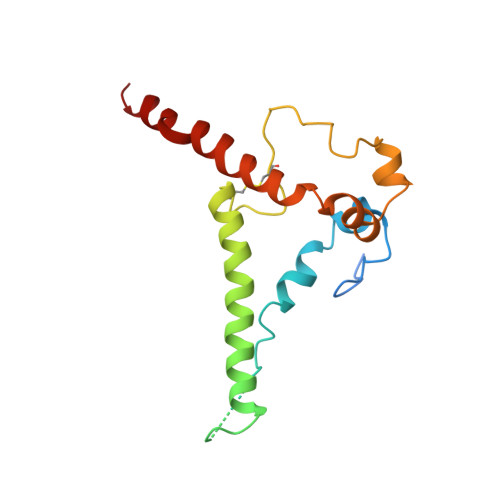
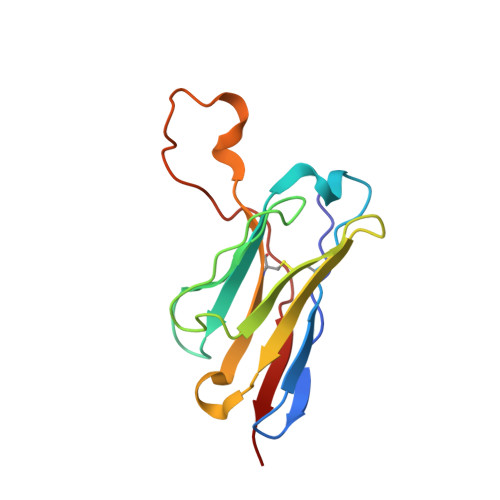
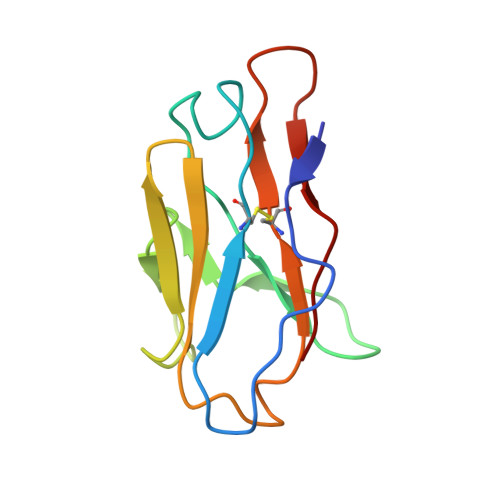
Stimulating broadly neutralizing antibodies (bnAbs) directly from germline remains a barrier for HIV vaccines. HIV superinfection elicits bnAbs more frequently than single infection, providing clues of how to elicit such responses. We used longitudinal antibody sequencing and structural studies to characterize bnAb development from a superinfection case. BnAb QA013.2 bound initial and superinfecting viral Env, despite its probable naive progenitor only recognizing the superinfecting strain, suggesting both viruses influenced this lineage. A 4.15 Å cryo-EM structure of QA013.2 bound to native-like trimer showed recognition of V3 signatures (N301/N332 and GDIR). QA013.2 relies less on CDRH3 and more on framework and CDRH1 for affinity and breadth compared to other V3/glycan-specific bnAbs. Antigenic profiling revealed that viral escape was achieved by changes in the structurally-defined epitope and by mutations in V1. These results highlight shared and novel properties of QA013.2 relative to other V3/glycan-specific bnAbs in the setting of sequential, diverse antigens.
- Division of Human Biology, Fred Hutchinson Cancer Research Center, Seattle, United States.
Organizational Affiliation: