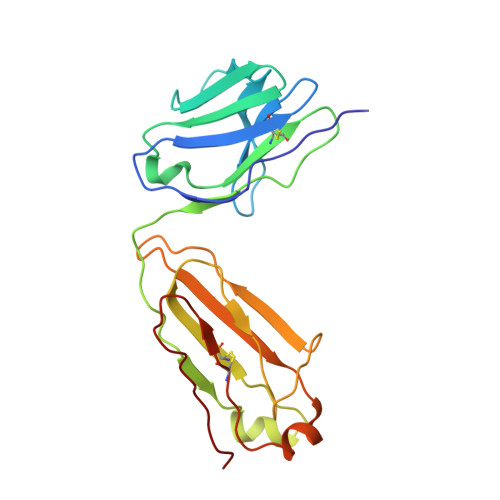
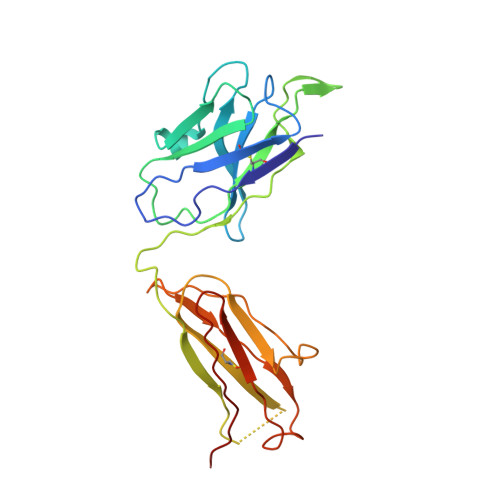
Development of Neutralization Breadth against Diverse HIV-1 by Increasing Ab-Ag Interface on V2.
Gao, N., Gai, Y., Meng, L., Wang, C., Wang, W., Li, X., Gu, T., Louder, M.K., Doria-Rose, N.A., Wiehe, K., Nazzari, A.F., Olia, A.S., Gorman, J., Rawi, R., Wu, W., Smith, C., Khant, H., de Val, N., Yu, B., Luo, J., Niu, H., Tsybovsky, Y., Liao, H., Kepler, T.B., Kwong, P.D., Mascola, J.R., Qin, C., Zhou, T., Yu, X., Gao, F.(2022) Adv Sci (Weinh) 9: e2200063-e2200063
- PubMed: 35319830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/advs.202200063
- Primary Citation Related Structures:
7MXD, 7N28 - PubMed Abstract:
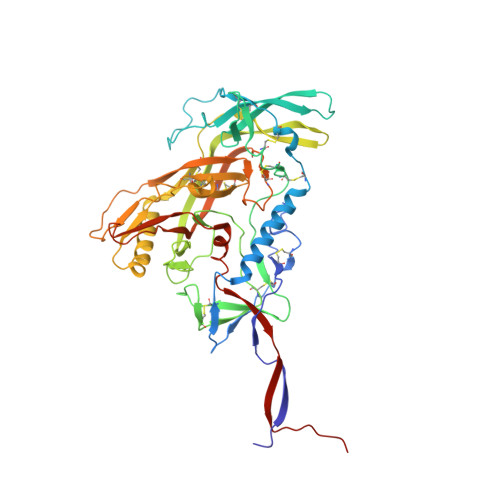
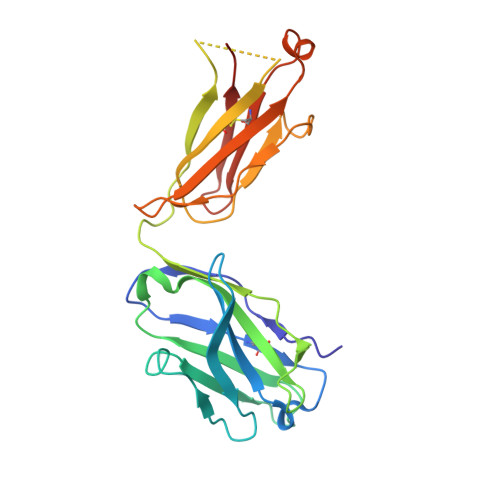
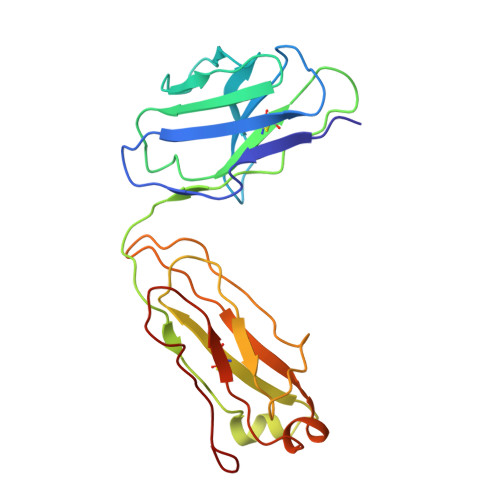
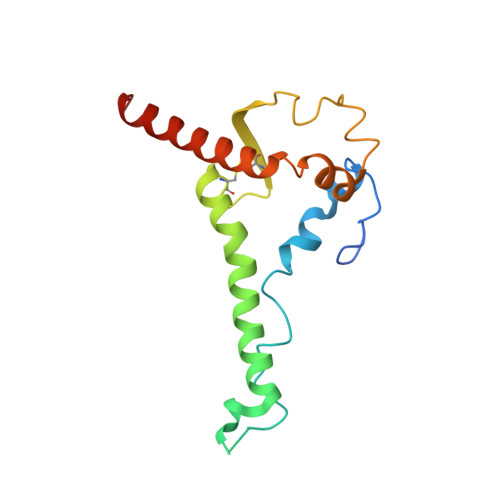
Understanding maturation pathways of broadly neutralizing antibodies (bnAbs) against HIV-1 can be highly informative for HIV-1 vaccine development. A lineage of J038 bnAbs is now obtained from a long-term SHIV-infected macaque. J038 neutralizes 54% of global circulating HIV-1 strains. Its binding induces a unique "up" conformation for one of the V2 loops in the trimeric envelope glycoprotein and is heavily dependent on glycan, which provides nearly half of the binding surface. Their unmutated common ancestor neutralizes the autologous virus. Continuous maturation enhances neutralization potency and breadth of J038 lineage antibodies via expanding antibody-Env contact areas surrounding the core region contacted by germline-encoded residues. Developmental details and recognition features of J038 lineage antibodies revealed here provide a new pathway for elicitation and maturation of V2-targeting bnAbs.
- National Engineering Laboratory for AIDS Vaccine, School of Life Sciences, Jilin University, Changchun, Jilin Province, 130012, China.
Organizational Affiliation: