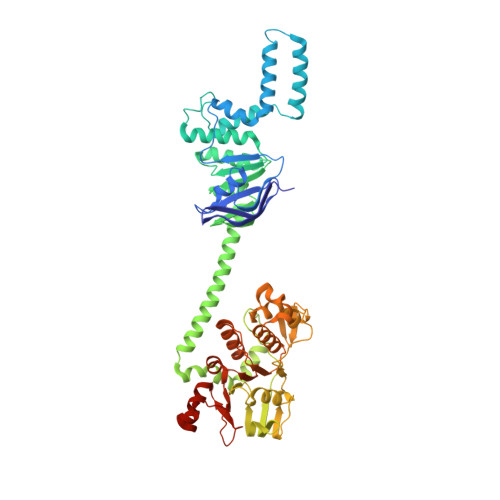
Interplay between an ATP-binding cassette F protein and the ribosome from Mycobacterium tuberculosis.
Cui, Z., Li, X., Shin, J., Gamper, H., Hou, Y.M., Sacchettini, J.C., Zhang, J.(2022) Nat Commun 13: 432-432
- PubMed: 35064151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28078-1
- Primary Citation Related Structures:
7MSC, 7MSH, 7MSM, 7MSZ, 7MT2, 7MT3, 7MT7, 7MU0 - PubMed Abstract:
EttA, energy-dependent translational throttle A, is a ribosomal factor that gates ribosome entry into the translation elongation cycle. A detailed understanding of its mechanism of action is limited due to the lack of high-resolution structures along its ATPase cycle. Here we present the cryo-electron microscopy (cryo-EM) structures of EttA from Mycobacterium tuberculosis (Mtb), referred to as MtbEttA, in complex with the Mtb 70S ribosome initiation complex (70SIC) at the pre-hydrolysis (ADPNP) and transition (ADP-VO 4 ) states, and the crystal structure of MtbEttA alone in the post-hydrolysis (ADP) state. We observe that MtbEttA binds the E-site of the Mtb 70SIC, remodeling the P-site tRNA and the ribosomal intersubunit bridge B7a during the ribosomal ratcheting. In return, the rotation of the 30S causes conformational changes in MtbEttA, forcing the two nucleotide-binding sites (NBSs) to alternate to engage each ADPNP in the pre-hydrolysis states, followed by complete engagements of both ADP-VO 4 molecules in the ATP-hydrolysis transition states. In the post-hydrolysis state, the conserved ATP-hydrolysis motifs of MtbEttA dissociate from both ADP molecules, leaving two nucleotide-binding domains (NBDs) in an open conformation. These structures reveal a dynamic interplay between MtbEttA and the Mtb ribosome, providing insights into the mechanism of translational regulation by EttA-like proteins.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX, 77843, USA.
Organizational Affiliation: