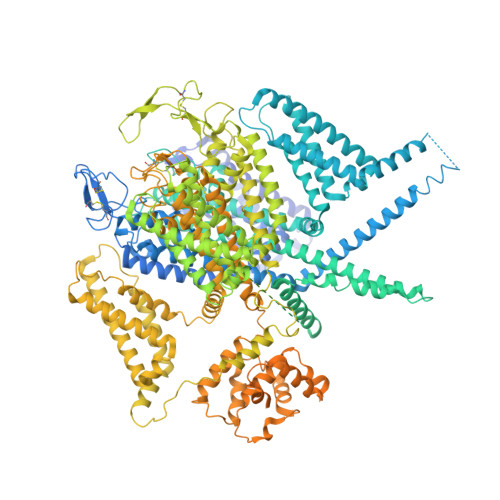
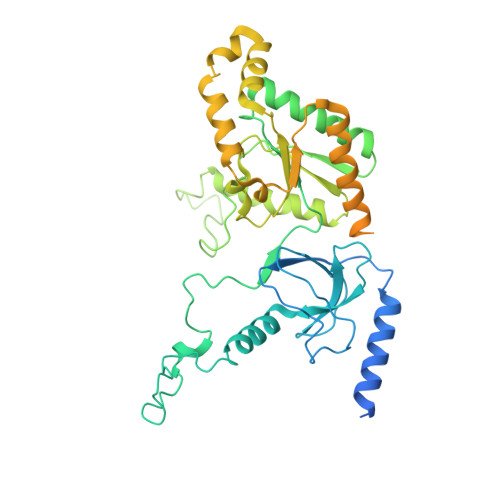
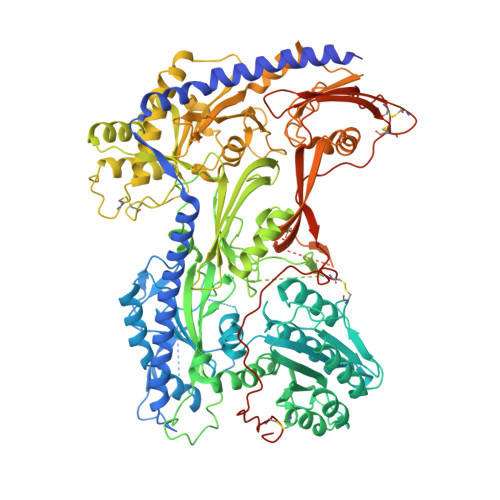
Structure of human Ca v 2.2 channel blocked by the painkiller ziconotide.
Gao, S., Yao, X., Yan, N.(2021) Nature 596: 143-147
- PubMed: 34234349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03699-6
- Primary Citation Related Structures:
7MIX, 7MIY - PubMed Abstract:
The neuronal-type (N-type) voltage-gated calcium (Ca v ) channels, which are designated Ca v 2.2, have an important role in the release of neurotransmitters 1-3 . Ziconotide is a Ca v 2.2-specific peptide pore blocker that has been clinically used for treating intractable pain 4-6 . Here we present cryo-electron microscopy structures of human Ca v 2.2 (comprising the core α1 and the ancillary α2δ-1 and β3 subunits) in the presence or absence of ziconotide. Ziconotide is thoroughly coordinated by helices P1 and P2, which support the selectivity filter, and the extracellular loops (ECLs) in repeats II, III and IV of α1. To accommodate ziconotide, the ECL of repeat III and α2δ-1 have to tilt upward concertedly. Three of the voltage-sensing domains (VSDs) are in a depolarized state, whereas the VSD of repeat II exhibits a down conformation that is stabilized by Ca v 2-unique intracellular segments and a phosphatidylinositol 4,5-bisphosphate molecule. Our studies reveal the molecular basis for Ca v 2.2-specific pore blocking by ziconotide and establish the framework for investigating electromechanical coupling in Ca v channels.
- Department of Molecular Biology, Princeton University, Princeton, NJ, USA.
Organizational Affiliation: