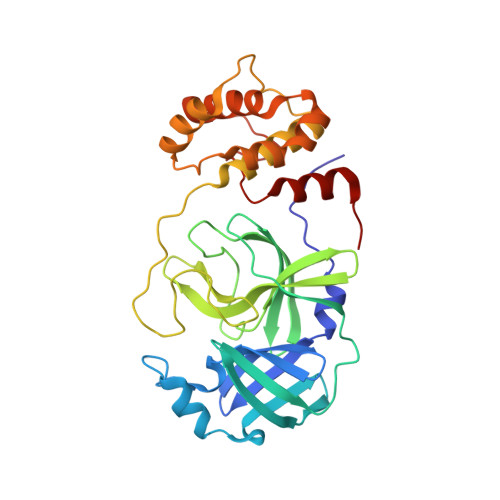
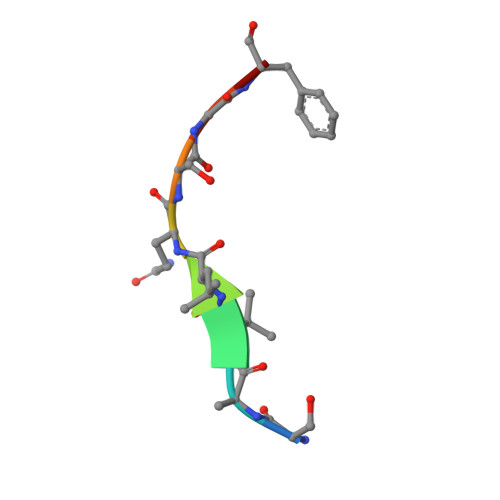
Recognition of Divergent Viral Substrates by the SARS-CoV-2 Main Protease.
MacDonald, E.A., Frey, G., Namchuk, M.N., Harrison, S.C., Hinshaw, S.M., Windsor, I.W.(2021) ACS Infect Dis 7: 2591-2595
- PubMed: 34437808 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.1c00237
- Primary Citation Related Structures:
7MGR, 7MGS - PubMed Abstract:
The main protease (M pro ) of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the cause of coronavirus disease (COVID-19), is an ideal target for pharmaceutical inhibition. M pro is conserved among coronaviruses and distinct from human proteases. Viral replication depends on the cleavage of the viral polyprotein at multiple sites. We present crystal structures of SARS-CoV-2 M pro bound to two viral substrate peptides. The structures show how M pro recognizes distinct substrates and how subtle changes in substrate accommodation can drive large changes in catalytic efficiency. One peptide, constituting the junction between viral nonstructural proteins 8 and 9 (nsp8/9), has P1' and P2' residues that are unique among the SARS-CoV-2 M pro cleavage sites but conserved among homologous junctions in coronaviruses. M pro cleaves nsp8/9 inefficiently, and amino acid substitutions at P1' or P2' can enhance catalysis. Visualization of M pro with intact substrates provides new templates for antiviral drug design and suggests that the coronavirus lifecycle selects for finely tuned substrate-dependent catalytic parameters.
- Laboratory of Molecular Medicine, Boston Children's Hospital, Boston, Massachusetts 02115, United States.
Organizational Affiliation: