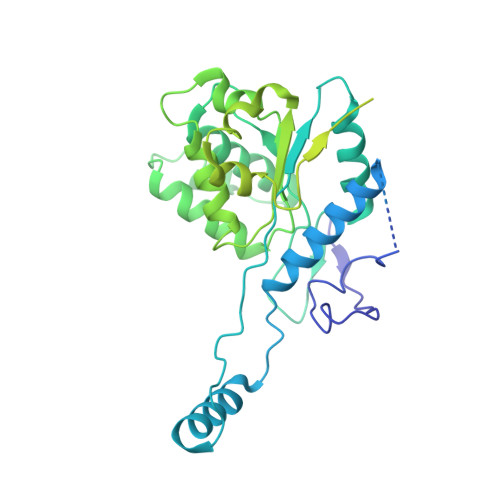
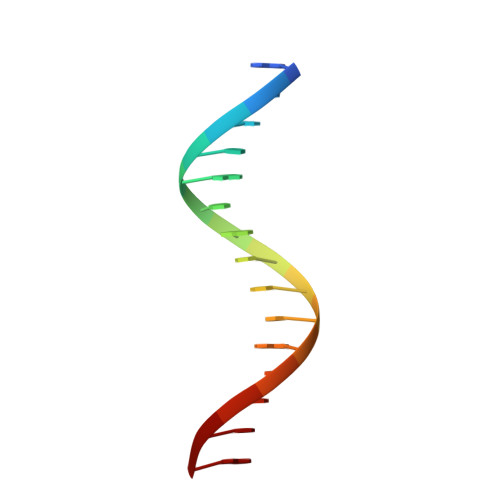
Structural basis for DNA targeting by the Tn7 transposon.
Shen, Y., Gomez-Blanco, J., Petassi, M.T., Peters, J.E., Ortega, J., Guarne, A.(2022) Nat Struct Mol Biol 29: 143-151
- PubMed: 35173349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00724-8
- Primary Citation Related Structures:
7MBW, 7MCS - PubMed Abstract:
Tn7 transposable elements are unique for their highly specific, and sometimes programmable, target-site selection mechanisms and precise insertions. All the elements in the Tn7 family utilize an AAA+ adaptor (TnsC) to coordinate target-site selection with transpososome assembly and to prevent insertions at sites already containing a Tn7 element. Owing to its multiple functions, TnsC is considered the linchpin in the Tn7 element. Here we present the high-resolution cryo-EM structure of TnsC bound to DNA using a gain-of-function variant of the protein and a DNA substrate that together recapitulate the recruitment to a specific DNA target site. TnsC forms an asymmetric ring on target DNA that segregates target-site selection and interaction with the paired-end complex to opposite faces of the ring. Unlike most AAA+ ATPases, TnsC uses a DNA distortion to find the target site but does not remodel DNA to activate transposition. By recognizing pre-distorted substrates, TnsC creates a built-in regulatory mechanism where ATP hydrolysis abolishes ring formation proximal to an existing element. This work unveils how Tn7 and Tn7-like elements determine the strict spacing between the target and integration sites.
- Department of Biochemistry, McGill University, Montreal, Quebec, Canada.
Organizational Affiliation: