Soluble Methane Monooxygenase Component Interactions Monitored by 19 F NMR.
Jones, J.C., Banerjee, R., Shi, K., Semonis, M.M., Aihara, H., Pomerantz, W.C.K., Lipscomb, J.D.(2021) Biochemistry 60: 1995-2010
- PubMed: 34100595 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.1c00293
- Primary Citation Related Structures:
7M8Q, 7M8R - PubMed Abstract:
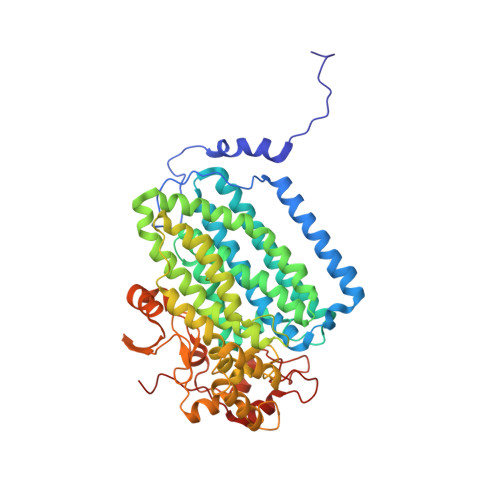
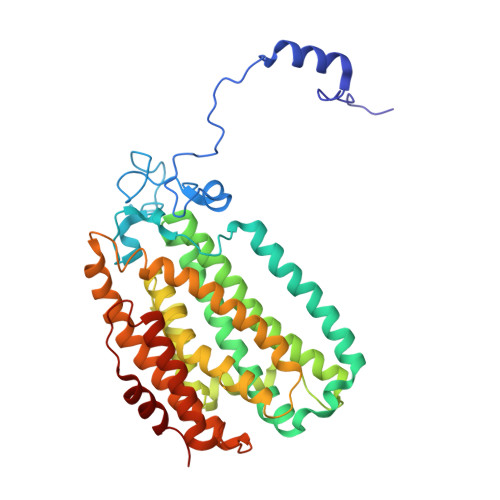
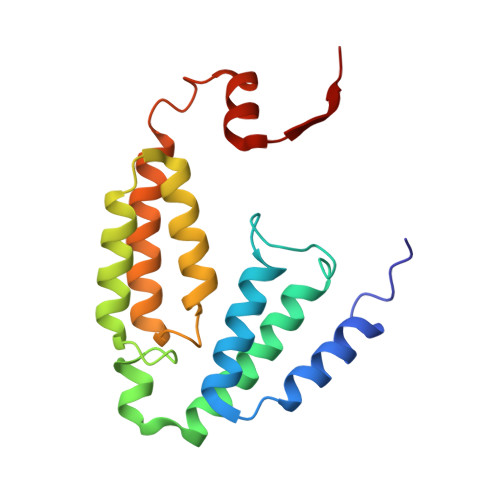
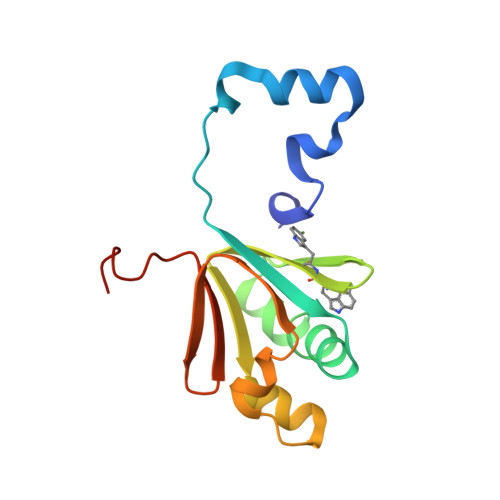
Soluble methane monooxygenase (sMMO) is a multicomponent metalloenzyme capable of catalyzing the fissure of the C-H bond of methane and the insertion of one atom of oxygen from O 2 to yield methanol. Efficient multiple-turnover catalysis occurs only in the presence of all three sMMO protein components: hydroxylase (MMOH), reductase (MMOR), and regulatory protein (MMOB). The complex series of sMMO protein component interactions that regulate the formation and decay of sMMO reaction cycle intermediates is not fully understood. Here, the two tryptophan residues in MMOB and the single tryptophan residue in MMOR are converted to 5-fluorotryptophan (5FW) by expression in defined media containing 5-fluoroindole. In addition, the mechanistically significant N-terminal region of MMOB is 19 F-labeled by reaction of the K15C variant with 3-bromo-1,1,1-trifluoroacetone (BTFA). The 5FW and BTFA modifications cause minimal structural perturbation, allowing detailed studies of the interactions with sMMOH using 19 F NMR. Resonances from the 275 kDa complexes of sMMOH with 5FW-MMOB and BTFA-K15C-5FW-MMOB are readily detected at 5 μM labeled protein concentration. This approach shows directly that MMOR and MMOB competitively bind to sMMOH with similar K D values, independent of the oxidation state of the sMMOH diiron cluster. These findings suggest a new model for regulation in which the dynamic equilibration of MMOR and MMOB with sMMOH allows a transient formation of key reactive complexes that irreversibly pull the reaction cycle forward. The slow kinetics of exchange of the sMMOH:MMOB complex is proposed to prevent MMOR-mediated reductive quenching of the high-valent reaction cycle intermediate Q before it can react with methane.