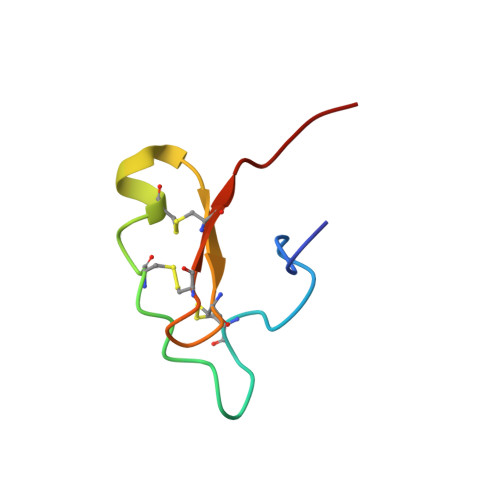
Structural and functional studies of scorpine: A channel blocker and cytolytic peptide.
Lopez-Giraldo, E., Carrillo, E., Titaux-Delgado, G., Cano-Sanchez, P., Colorado, A., Possani, L.D., Rio-Portilla, F.D.(2022) Toxicon 222: 106985-106985
- PubMed: 36436588 Search on PubMed
- DOI: https://doi.org/10.1016/j.toxicon.2022.106985
- Primary Citation Related Structures:
7M1D, 7M1E - PubMed Abstract:
Scorpine is an antimicrobial and antimalarial peptide isolated from Pandinus imperator scorpion venom. As there are few functional and structural studies reported on scorpine-like peptides, we investigated the recombinant truncated N- and C-terminal domains as well as complete scorpine using biological assays and determined the N- and C-terminal structures using solution nuclear magnetic resonance. The study was conducted using recombinant N- and C-terminal peptides and complete scorpine expressed in Escherichia coli. The results showed that N-scorpine presented a random coil structure in water and adopted α-helical folding in the presence of 50% trifluoroethanol (TFE). C-scorpine contains three disulfide bonds with two structural domains: an unstructured N-terminal domain in water that can form a typical secondary alpha-helix structure in 50% TFE and a C-terminal domain with the CS-αβ motif. Our findings demonstrate cytolytic activity associated with C-scorpine, N-scorpine, and scorpine, as well as channel blocking activity associated with the C-scorpine domain.
- Instituto de Química, Universidad Nacional Autónoma de México, CdMx, Mexico.
Organizational Affiliation: