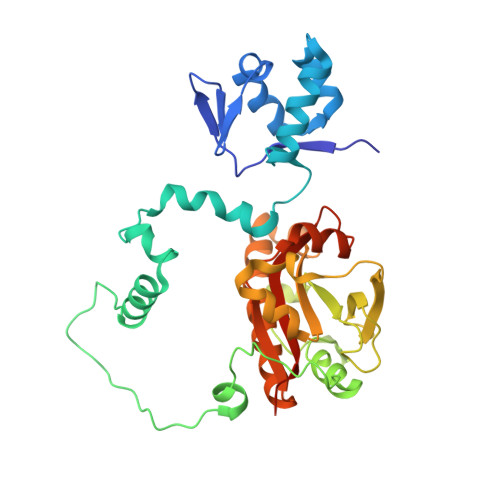
Structures and function of a tailoring oxidase in complex with a nonribosomal peptide synthetase module.
Fortinez, C.M., Bloudoff, K., Harrigan, C., Sharon, I., Strauss, M., Schmeing, T.M.(2022) Nat Commun 13: 548-548
- PubMed: 35087027 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28221-y
- Primary Citation Related Structures:
7LY4, 7LY5, 7LY6, 7LY7 - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are large modular enzymes that synthesize secondary metabolites and natural product therapeutics. Most NRPS biosynthetic pathways include an NRPS and additional proteins that introduce chemical modifications before, during or after assembly-line synthesis. The bacillamide biosynthetic pathway is a common, three-protein system, with a decarboxylase that prepares an NRPS substrate, an NRPS, and an oxidase. Here, the pathway is reconstituted in vitro. The oxidase is shown to perform dehydrogenation of the thiazoline in the peptide intermediate while it is covalently attached to the NRPS, as the penultimate step in bacillamide D synthesis. Structural analysis of the oxidase reveals a dimeric, two-lobed architecture with a remnant RiPP recognition element and a dramatic wrapping loop. The oxidase forms a stable complex with the NRPS and dimerizes it. We visualized co-complexes of the oxidase bound to the elongation module of the NRPS using X-ray crystallography and cryo-EM. The three active sites (for adenylation, condensation/cyclization, and oxidation) form an elegant arc to facilitate substrate delivery. The structures enabled a proof-of-principle bioengineering experiment in which the BmdC oxidase domain is embedded into the NRPS.
- Department of Biochemistry, McGill University, Montréal, QC, H3G 0B1, Canada.
Organizational Affiliation: