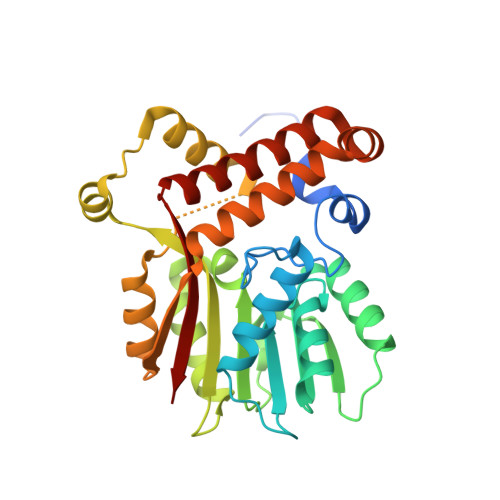
Crystal structure of the S-adenosylmethionine-dependent mycolic acid synthase UmaA from Mycobacterium tuberculosis.
Teng, S., Wang, J., Sroge, C.D., Abendroth, J., Lorimer, D.D., Horanyi, P.S., Edwards, T.E., Tillery, L., Craig, J.K., Van Voorhis, W.C., Myler, P.J., Smith, C.L.(2025) Acta Crystallogr F Struct Biol Commun 81: 146-154
- PubMed: 40059638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25001530
- Primary Citation Related Structures:
7LXI - PubMed Abstract:
Mycobacterium tuberculosis is a Gram-positive bacillus that causes tuberculosis and is a leading cause of mortality worldwide. This disease is a growing health threat due to the occurrence of multidrug resistance. Mycolic acids are essential for generating cell walls and their modification is important to the virulence and persistence of M. tuberculosis. A family of S-adenosylmethionine-dependent mycolic acid synthases modify mycolic acids and represent promising drug targets. UmaA is currently the least-understood member of this family. This paper describes the crystal structure of UmaA. UmaA is a monomer composed of two domains: a structurally conserved SAM-binding domain and a variable substrate-binding auxiliary domain. Fortuitously, our structure contains a nitrate in the active site, a structural mimic of carbonate, which is a known general base in cyclopropane-adding synthases. Further investigation indicated that the structure of the N-terminus is highly flexible. Finally, we have identified S-adenosyl-N-decyl-aminoethyl as a promising potential inhibitor.
- Department of Biology, Washington University in St Louis, St Louis, MO 63134, USA.
Organizational Affiliation: