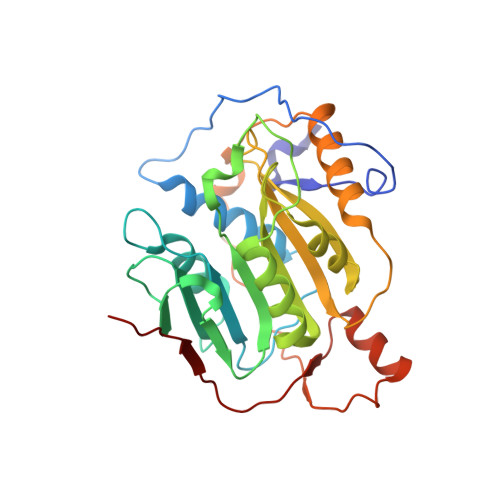
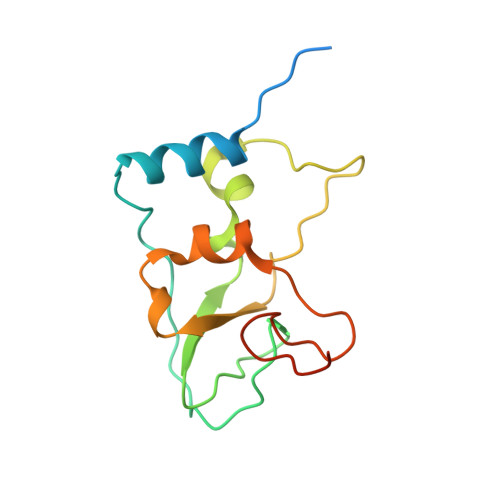
A metal ion orients SARS-CoV-2 mRNA to ensure accurate 2'-O methylation of its first nucleotide.
Viswanathan, T., Misra, A., Chan, S.H., Qi, S., Dai, N., Arya, S., Martinez-Sobrido, L., Gupta, Y.K.(2021) Nat Commun 12: 3287-3287
- PubMed: 34078893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-23594-y
- Primary Citation Related Structures:
7LW3, 7LW4 - PubMed Abstract:
The SARS-CoV-2 nsp16/nsp10 enzyme complex modifies the 2'-OH of the first transcribed nucleotide of the viral mRNA by covalently attaching a methyl group to it. The 2'-O methylation of the first nucleotide converts the status of mRNA cap from Cap-0 to Cap-1, and thus, helps the virus evade immune surveillance in host cells. Here, we report two structures of nsp16/nsp10 representing pre- and post-release states of the RNA product (Cap-1). We observe overall widening of the enzyme upon product formation, and an inward twisting motion in the substrate binding region upon product release. These conformational changes reset the enzyme for the next round of catalysis. The structures also identify a unique binding mode and the importance of a divalent metal ion for 2'-O methylation. We also describe underlying structural basis for the perturbed enzymatic activity of a clinical variant of SARS-CoV-2, and a previous SARS-CoV outbreak strain.
- Greehey Children's Cancer Research Institute, University of Texas Health at San Antonio, San Antonio, TX, USA.
Organizational Affiliation: