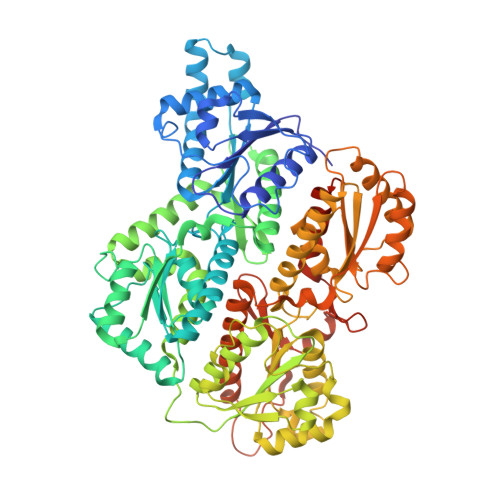
Selective activation of PFKL suppresses the phagocytic oxidative burst.
Amara, N., Cooper, M.P., Voronkova, M.A., Webb, B.A., Lynch, E.M., Kollman, J.M., Ma, T., Yu, K., Lai, Z., Sangaraju, D., Kayagaki, N., Newton, K., Bogyo, M., Staben, S.T., Dixit, V.M.(2021) Cell 184: 4480-4494.e15
- PubMed: 34320407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2021.07.004
- Primary Citation Related Structures:
7LW1 - PubMed Abstract:
In neutrophils, nicotinamide adenine dinucleotide phosphate (NADPH) generated via the pentose phosphate pathway fuels NADPH oxidase NOX2 to produce reactive oxygen species for killing invading pathogens. However, excessive NOX2 activity can exacerbate inflammation, as in acute respiratory distress syndrome (ARDS). Here, we use two unbiased chemical proteomic strategies to show that small-molecule LDC7559, or a more potent designed analog NA-11, inhibits the NOX2-dependent oxidative burst in neutrophils by activating the glycolytic enzyme phosphofructokinase-1 liver type (PFKL) and dampening flux through the pentose phosphate pathway. Accordingly, neutrophils treated with NA-11 had reduced NOX2-dependent outputs, including neutrophil cell death (NETosis) and tissue damage. A high-resolution structure of PFKL confirmed binding of NA-11 to the AMP/ADP allosteric activation site and explained why NA-11 failed to agonize phosphofructokinase-1 platelet type (PFKP) or muscle type (PFKM). Thus, NA-11 represents a tool for selective activation of PFKL, the main phosphofructokinase-1 isoform expressed in immune cells.
- Physiological Chemistry Department, Genentech, South San Francisco, CA 94080, USA.
Organizational Affiliation: