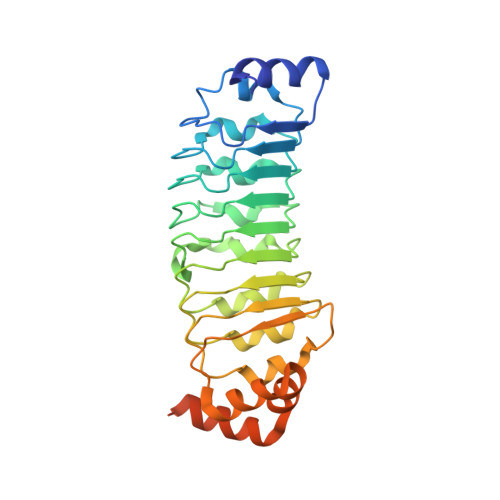
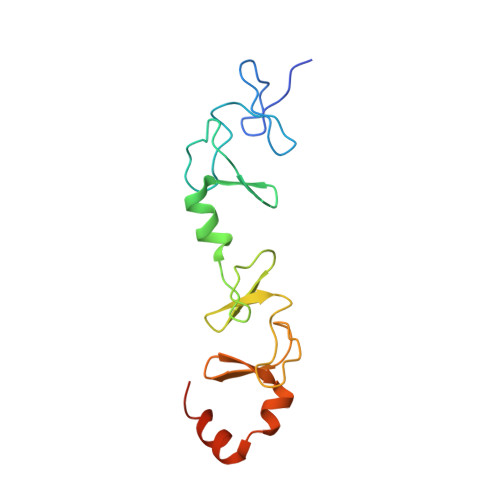
Molecular basis for Ras suppressor-1 binding to PINCH-1 in focal adhesion assembly.
Fukuda, K., Lu, F., Qin, J.(2021) J Biological Chem 296: 100685-100685
- PubMed: 33891945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100685
- Primary Citation Related Structures:
7LT8, 7LT9 - PubMed Abstract:
Ras suppressor-1 (Rsu-1) is a leucine-rich repeat (LRR)-containing protein that is crucial for regulating cell adhesion and is involved in such physiological and pathological processes as focal adhesion assembly and tumor metastasis. Rsu-1 interacts with zinc-finger type multi-LIM domain-containing adaptor protein PINCH-1, known to be involved in the integrin-mediated consensus adhesome, but not with its highly homologous family member PINCH-2. However, the structural basis for and regulatory mechanisms of this specific interaction remain unclear. Here, we determined the crystal structures of Rsu-1 and its complex with the PINCH-1 LIM4-5 domains. Rsu-1 displays an arc-shaped solenoid architecture, with eight LRRs shielded by N- and C-terminal capping modules. We showed that the conserved concave surface of the Rsu-1 LRR domain binds and stabilizes the PINCH-1 LIM5 domain via salt bridge and hydrophobic interactions, while the C-terminal non-LIM region of PINCH-2 sterically disfavors Rsu-1 binding. We also showed that Rsu-1 can be assembled, via PINCH-1-binding, into a heteropentamer complex comprising Rsu-1, PINCH-1, ILK, Parvin, and Kindlin-2, which constitute a major consensus integrin adhesome crucial for focal adhesion assembly. Our mutagenesis and cell biological data emphasize the significance of the Rsu-1/PINCH-1 interaction in focal adhesion assembly and cell spreading, providing crucial molecular insights into Rsu-1-mediated cell adhesion with implications for disease development.
- Department of Cardiovascular & Metabolic Sciences, Lerner Research Institute, Cleveland Clinic, Ohio, USA.
Organizational Affiliation: