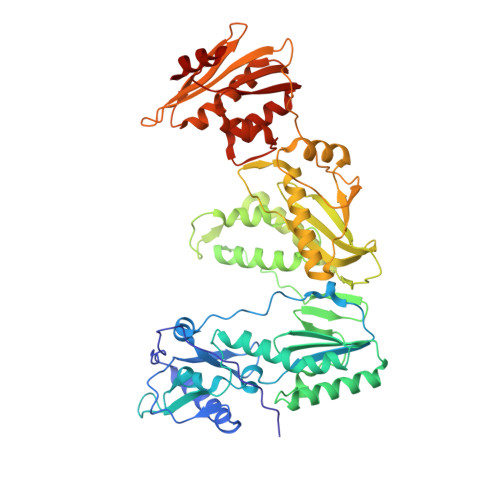
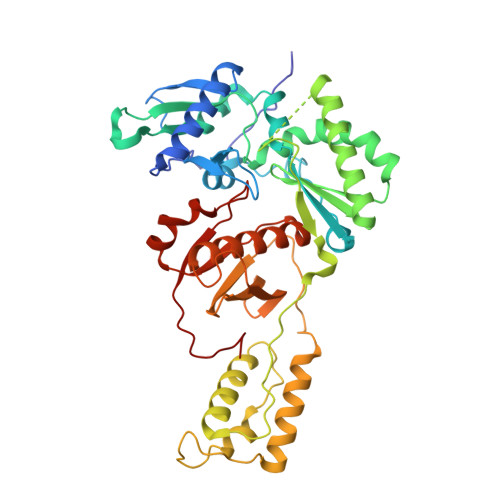
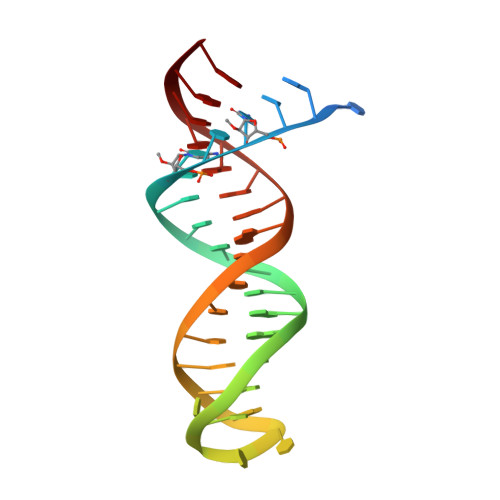
Structural basis of HIV inhibition by L-nucleosides: Opportunities for drug development and repurposing.
Ruiz, F.X., Hoang, A., Dilmore, C.R., DeStefano, J.J., Arnold, E.(2022) Drug Discov Today 27: 1832-1846
- PubMed: 35218925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.drudis.2022.02.016
- Primary Citation Related Structures:
7LRI, 7LRM, 7LRX, 7LRY, 7LSK - PubMed Abstract:
Infection with HIV can cripple the immune system and lead to AIDS. Hepatitis B virus (HBV) is a hepadnavirus that causes human liver diseases. Both pathogens are major public health problems affecting millions of people worldwide. The polymerases from both viruses are the most common drug target for viral inhibition, sharing common architecture at their active sites. The L-nucleoside drugs emtricitabine and lamivudine are widely used HIV reverse transcriptase (RT) and HBV polymerase (Pol) inhibitors. Nevertheless, structural details of their binding to RT(Pol)/nucleic acid remained unknown until recently. Here, we discuss the implications of these structures, alongside related complexes with L-dNTPs, for the development of novel L-nucleos(t)ide drugs, and prospects for repurposing them.
- Center for Advanced Biotechnology and Medicine, and Department of Chemistry and Chemical Biology, Rutgers University, Piscataway, NJ 08854, USA. Electronic address: xavier@cabm.rutgers.edu.
Organizational Affiliation: