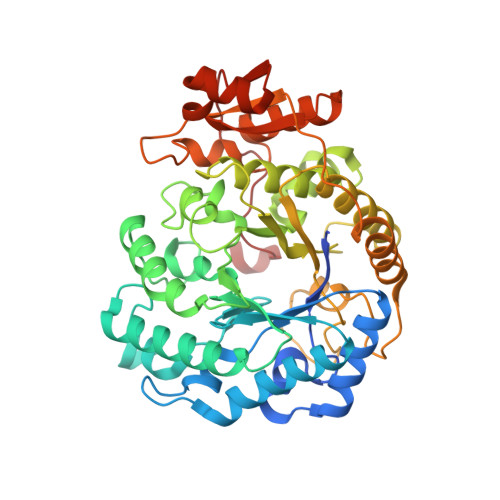
N-Glycan Degradation Pathways in Gut- and Soil-Dwelling Actinobacteria Share Common Core Genes.
Higgins, M.A., Tegl, G., MacDonald, S.S., Arnal, G., Brumer, H., Withers, S.G., Ryan, K.S.(2021) ACS Chem Biol 16: 701-711
- PubMed: 33764747 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.0c00995
- Primary Citation Related Structures:
7LQX, 7LR1, 7LR2, 7LR6, 7LR7, 7LR8 - PubMed Abstract:
N-Glycosylation is a fundamental protein modification found in both eukaryotes and archaea. Despite lacking N-glycans, many commensal and pathogenic bacteria have developed mechanisms to degrade these isoforms for a variety of functions, including nutrient acquisition and evasion of the immune system. Although much is known about many of the enzymes responsible for N-glycan degradation, the enzymes involved in cleaving the N-glycan core have only recently been discovered. Thus, some of the structural details have yet to be characterized, and little is known about their full distribution among bacterial strains and specifically within potential Gram-positive polysaccharide utilization loci. Here, we report crystal structures for Family 5, Subfamily 18 (GH5_18) glycoside hydrolases from the gut bacterium Bifidobacterium longum (BlGH5_18) and the soil bacterium Streptomyces cattleya (ScGH5_18), which hydrolyze the core Manβ1-4GlcNAc disaccharide. Structures of these enzymes in complex with Manβ1-4GlcNAc reveal a more complete picture of the -1 subsite. They also show that a C-terminal active site cap present in BlGH5_18 is absent in ScGH5_18. Although this C-terminal cap is not widely distributed throughout the GH5_18 family, it is important for full enzyme activity. In addition, we show that GH5_18 enzymes are found in Gram-positive polysaccharide utilization loci that share common genes, likely dedicated to importing and degrading N-glycan core structures.