Structural basis of malaria transmission blockade by a monoclonal antibody to gamete fusogen HAP2.
Feng, J., Dong, X., DeCosta, A., Su, Y., Angrisano, F., Sala, K.A., Blagborough, A.M., Lu, C., Springer, T.A.(2021) Elife 10
- PubMed: 34939934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.74707
- Primary Citation Related Structures:
7LR3, 7LR4 - PubMed Abstract:
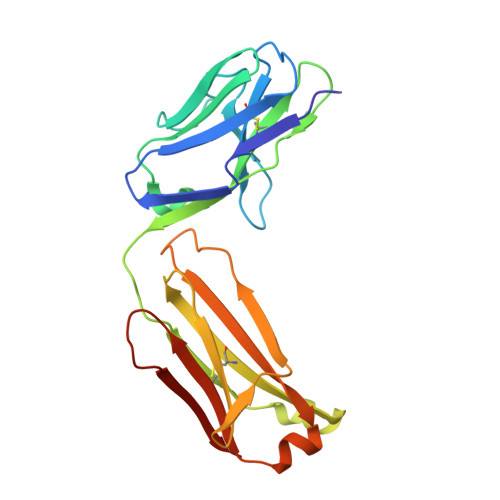
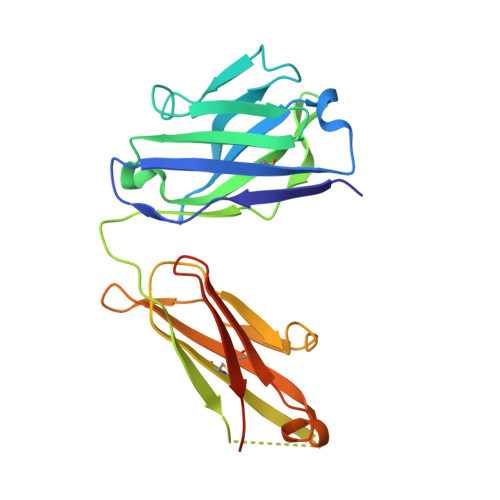
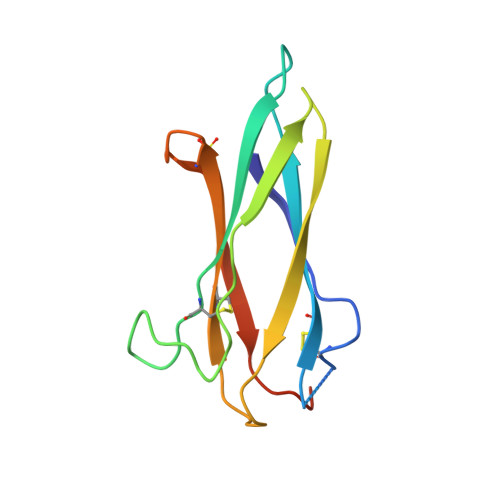
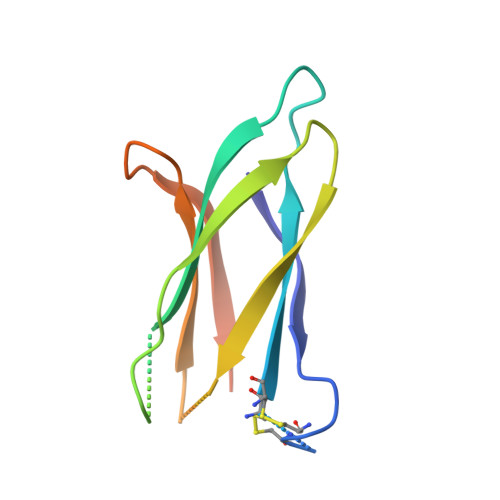
HAP2 is a transmembrane gamete fusogen found in multiple eukaryotic kingdoms and is structurally homologous to viral class II fusogens. Studies in Plasmodium have suggested that HAP2 is an attractive target for vaccines that block transmission of malaria. HAP2 has three extracellular domains, arranged in the order D2, D1, and D3. Here, we report monoclonal antibodies against the D3 fragment of Plasmodium berghei HAP2 and crystal structures of D3 in complex with Fab fragments of two of these antibodies, one of which blocks fertilization of Plasmodium berghei in vitro and transmission of malaria in mosquitoes. We also show how this Fab binds the complete HAP2 ectodomain with electron microscopy. The two antibodies cross-react with HAP2 among multiple plasmodial species. Our characterization of the Plasmodium D3 structure, HAP2 ectodomain architecture, and mechanism of inhibition provide insights for the development of a vaccine to block malaria transmission.
- Program in Cellular and Molecular Medicine, Boston Children's Hospital, Boston, United States.
Organizational Affiliation: