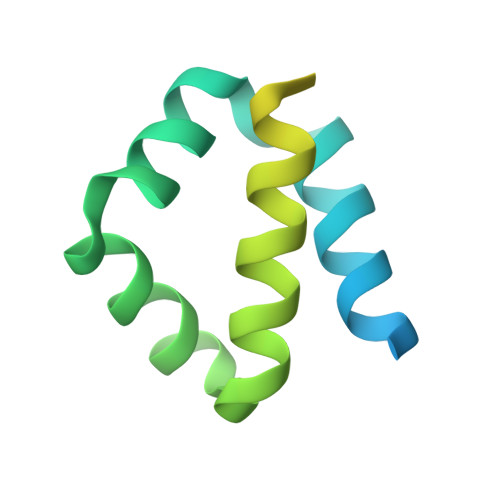
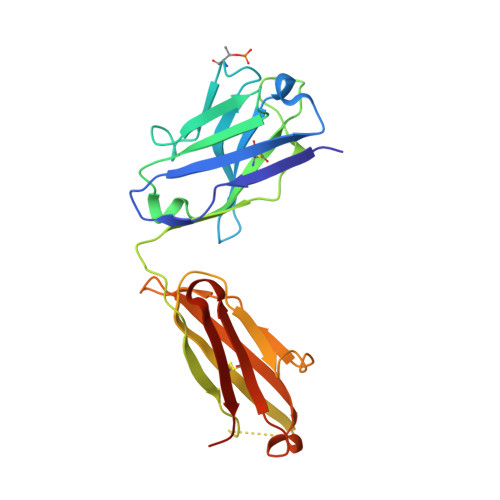
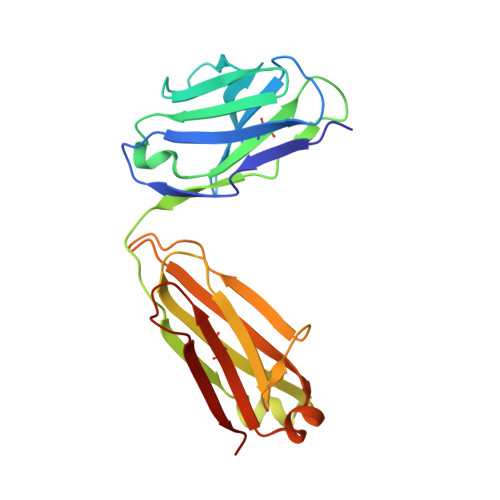
Structures of the ApoL1 and ApoL2 N-terminal domains reveal a non-classical four-helix bundle motif.
Ultsch, M., Holliday, M.J., Gerhardy, S., Moran, P., Scales, S.J., Gupta, N., Oltrabella, F., Chiu, C., Fairbrother, W., Eigenbrot, C., Kirchhofer, D.(2021) Commun Biol 4: 916-916
- PubMed: 34316015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-021-02387-5
- Primary Citation Related Structures:
7L6K, 7LF7, 7LF8, 7LFA, 7LFB, 7LFD - PubMed Abstract:
Apolipoprotein L1 (ApoL1) is a circulating innate immunity protein protecting against trypanosome infection. However, two ApoL1 coding variants are associated with a highly increased risk of chronic kidney disease. Here we present X-ray and NMR structures of the N-terminal domain (NTD) of ApoL1 and of its closest relative ApoL2. In both proteins, four of the five NTD helices form a four-helix core structure which is different from the classical four-helix bundle and from the pore-forming domain of colicin A. The reactivity with a conformation-specific antibody and structural models predict that this four-helix motif is also present in the NTDs of ApoL3 and ApoL4, suggesting related functions within the small ApoL family. The long helix 5 of ApoL1 is conformationally flexible and contains the BH3-like region. This BH3-like α-helix resembles true BH3 domains only in sequence and structure but not in function, since it does not bind to the pro-survival members of the Bcl-2 family, suggesting a Bcl-2-independent role in cytotoxicity. These findings should expedite a more comprehensive structural and functional understanding of the ApoL immune protein family.
- Department of Structural Biology, Genentech Inc., South San Francisco, CA, USA.
Organizational Affiliation: