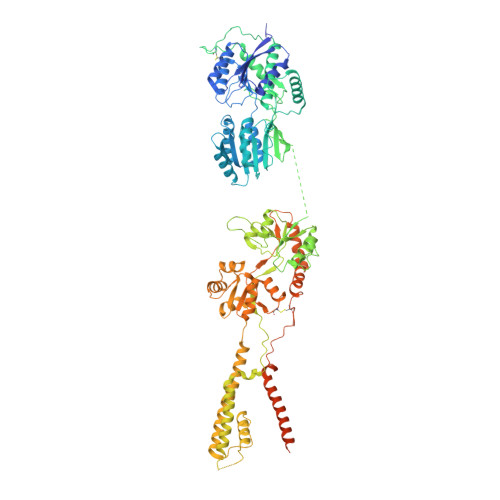
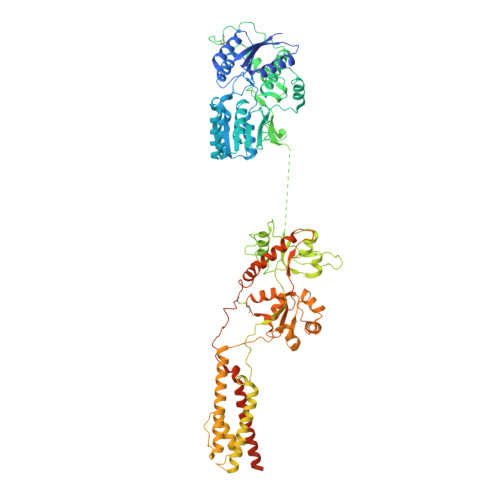
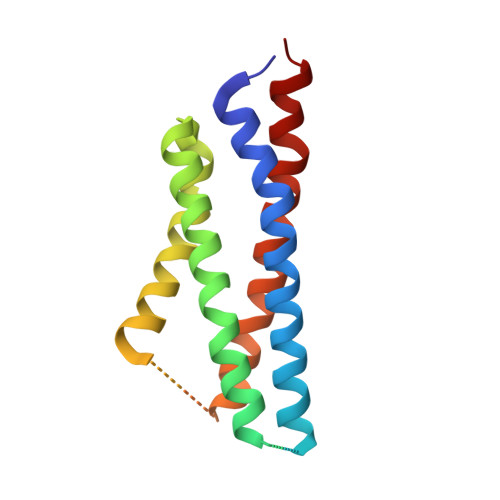
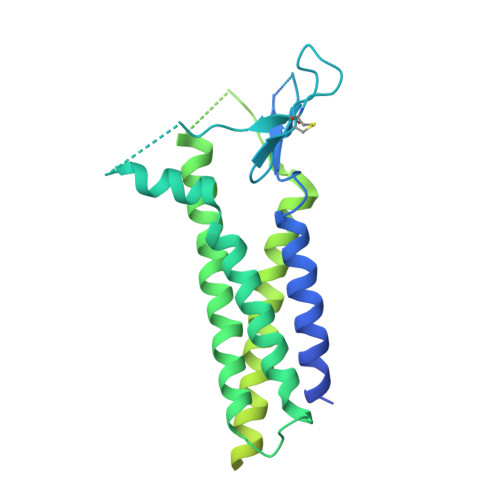
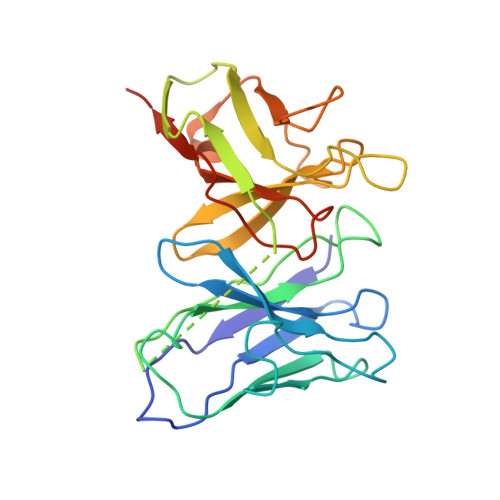
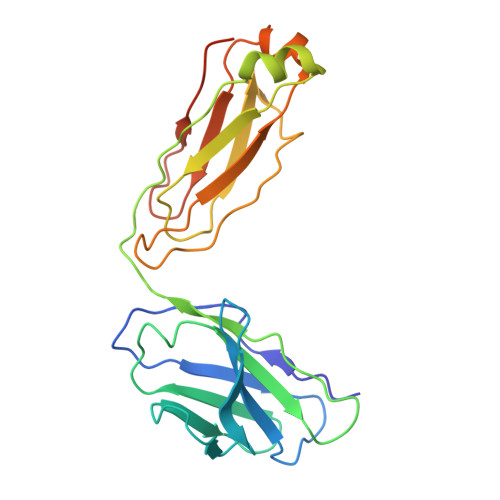
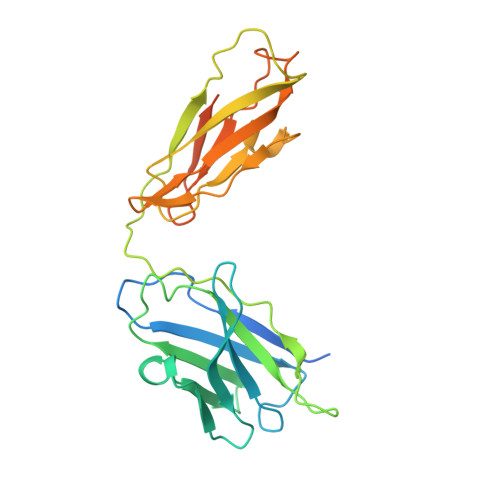
Hippocampal AMPA receptor assemblies and mechanism of allosteric inhibition.
Yu, J., Rao, P., Clark, S., Mitra, J., Ha, T., Gouaux, E.(2021) Nature 594: 448-453
- PubMed: 33981040 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03540-0
- Primary Citation Related Structures:
7LDD, 7LDE, 7LEP - PubMed Abstract:
AMPA-selective glutamate receptors mediate the transduction of signals between the neuronal circuits of the hippocampus 1 . The trafficking, localization, kinetics and pharmacology of AMPA receptors are tuned by an ensemble of auxiliary protein subunits, which are integral membrane proteins that associate with the receptor to yield bona fide receptor signalling complexes 2 . Thus far, extensive studies of recombinant AMPA receptor-auxiliary subunit complexes using engineered protein constructs have not been able to faithfully elucidate the molecular architecture of hippocampal AMPA receptor complexes. Here we obtain mouse hippocampal, calcium-impermeable AMPA receptor complexes using immunoaffinity purification and use single-molecule fluorescence and cryo-electron microscopy experiments to elucidate three major AMPA receptor-auxiliary subunit complexes. The GluA1-GluA2, GluA1-GluA2-GluA3 and GluA2-GluA3 receptors are the predominant assemblies, with the auxiliary subunits TARP-γ8 and CNIH2-SynDIG4 non-stochastically positioned at the B'/D' and A'/C' positions, respectively. We further demonstrate how the receptor-TARP-γ8 stoichiometry explains the mechanism of and submaximal inhibition by a clinically relevant, brain-region-specific allosteric inhibitor.
- Vollum Institute, Oregon Health & Science University, Portland, OR, USA.
Organizational Affiliation: