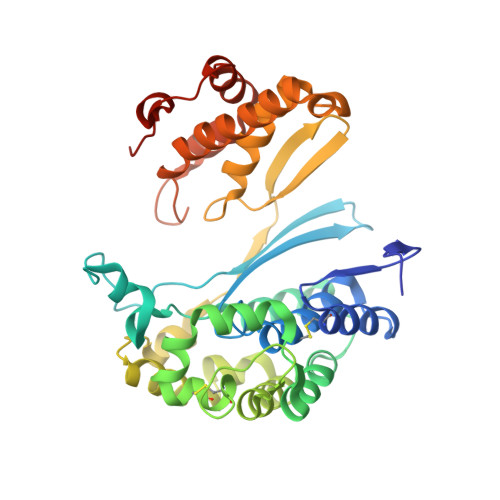
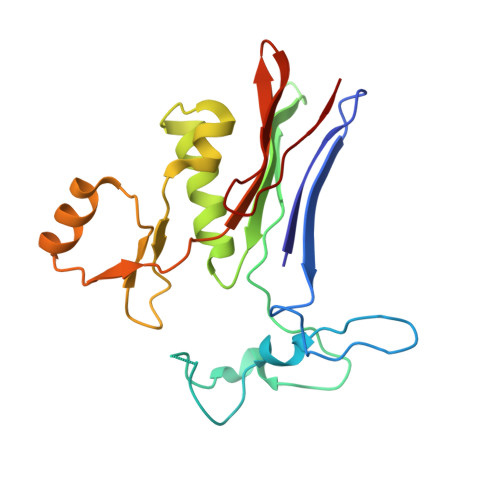
Design and evaluation of novel analogs of 2-amino-4-boronobutanoic acid (ABBA) as inhibitors of human gamma-glutamyl transpeptidase.
Nguyen, L., Schultz, D.C., Terzyan, S.S., Rezaei, M., Songb, J., Li, C., You, Y., Hanigan, M.H.(2022) Bioorg Med Chem 73: 116986-116986
- PubMed: 36208545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmc.2022.116986
- Primary Citation Related Structures:
7LBC, 7LD9 - PubMed Abstract:
Inhibitors of gamma-glutamyl transpeptidase (GGT1, aka gamma-glutamyl transferase) are needed for the treatment of cancer, cardiovascular illness and other diseases. Compounds that inhibit GGT1 have been evaluated in the clinic, but no inhibitor has successfully demonstrated specific and systemic GGT1 inhibition. All have severe side effects. L-2-amino-4‑boronobutanoic acid (l-ABBA), a glutamate analog, is the most potent GGT1 inhibitor in vitro. In this study, we have solved the crystal structure of human GGT1 (hGGT1) with ABBA bound in the active site. The structure was interrogated to identify interactions between the enzyme and the inhibitor. Based on these data, a series of novel ABBA analogs were designed and synthesized. Their inhibitory activity against the hydrolysis and transpeptidation activities of hGGT1 were determined. The lead compounds were crystalized with hGGT1 and the structures solved. The kinetic data and structures of the complexes provide new insights into the critical role of protein structure dynamics in developing compounds for inhibition of hGGT1.
- Department of Pharmaceutical Sciences, The State University of New York at Buffalo, Buffalo, NY 14214, United States.
Organizational Affiliation: