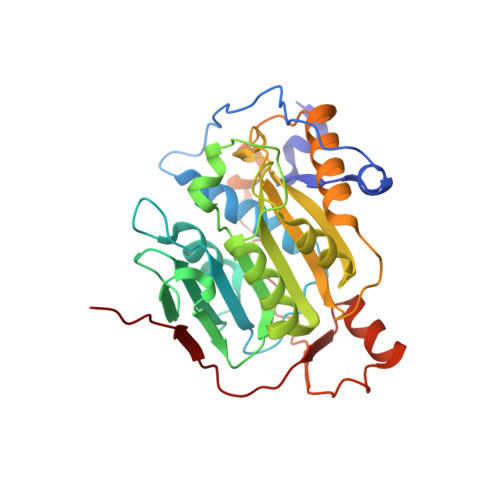
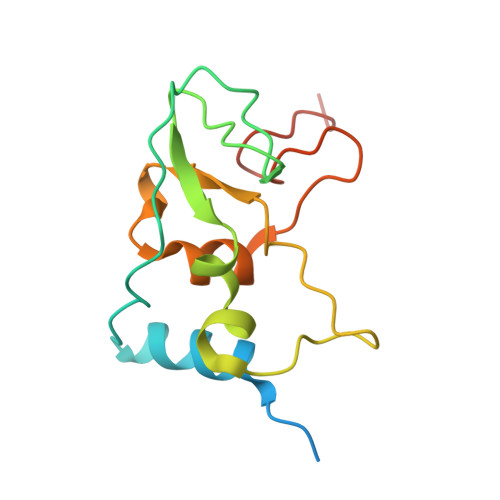
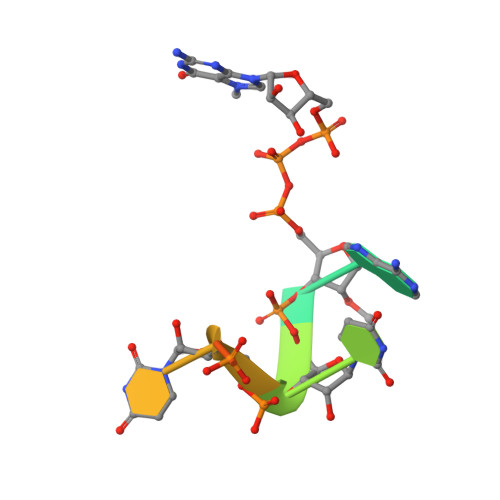
Mn 2+ coordinates Cap-0-RNA to align substrates for efficient 2'- O -methyl transfer by SARS-CoV-2 nsp16.
Minasov, G., Rosas-Lemus, M., Shuvalova, L., Inniss, N.L., Brunzelle, J.S., Daczkowski, C.M., Hoover, P., Mesecar, A.D., Satchell, K.J.F.(2021) Sci Signal 14
- PubMed: 34131072 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scisignal.abh2071
- Primary Citation Related Structures:
7JYY, 7L6R, 7L6T - PubMed Abstract:
Capping of viral messenger RNAs is essential for efficient translation, for virus replication, and for preventing detection by the host cell innate response system. The SARS-CoV-2 genome encodes the 2'- O -methyltransferase nsp16, which, when bound to the coactivator nsp10, uses S -adenosylmethionine (SAM) as a donor to transfer a methyl group to the first ribonucleotide of the mRNA in the final step of viral mRNA capping. Here, we provide biochemical and structural evidence that this reaction requires divalent cations, preferably Mn 2+ , and a coronavirus-specific four-residue insert. We determined the x-ray structures of the SARS-CoV-2 2'- O -methyltransferase (the nsp16-nsp10 heterodimer) in complex with its reaction substrates, products, and divalent metal cations. These structural snapshots revealed that metal ions and the insert stabilize interactions between the capped RNA and nsp16, resulting in the precise alignment of the ribonucleotides in the active site. Comparison of available structures of 2'- O -methyltransferases with capped RNAs from different organisms revealed that the four-residue insert unique to coronavirus nsp16 alters the backbone conformation of the capped RNA in the binding groove, thereby promoting catalysis. This insert is highly conserved across coronaviruses, and its absence in mammalian methyltransferases makes this region a promising site for structure-guided drug design of selective coronavirus inhibitors.
- Department of Microbiology-Immunology, Northwestern University, Feinberg School of Medicine,,Chicago, IL 60611, USA.
Organizational Affiliation: