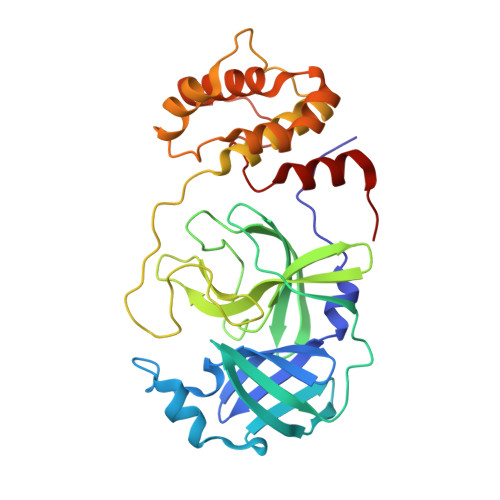
Crystal Structure of SARS-CoV-2 Main Protease in Complex with the Non-Covalent Inhibitor ML188.
Lockbaum, G.J., Reyes, A.C., Lee, J.M., Tilvawala, R., Nalivaika, E.A., Ali, A., Kurt Yilmaz, N., Thompson, P.R., Schiffer, C.A.(2021) Viruses 13
- PubMed: 33503819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/v13020174
- Primary Citation Related Structures:
7L0D - PubMed Abstract:
Viral proteases are critical enzymes for the maturation of many human pathogenic viruses and thus are key targets for direct acting antivirals (DAAs). The current viral pandemic caused by SARS-CoV-2 is in dire need of DAAs. The Main protease (M pro ) is the focus of extensive structure-based drug design efforts which are mostly covalent inhibitors targeting the catalytic cysteine. ML188 is a non-covalent inhibitor designed to target SARS-CoV-1 M pro , and provides an initial scaffold for the creation of effective pan-coronavirus inhibitors. In the current study, we found that ML188 inhibits SARS-CoV-2 M pro at 2.5 µM, which is more potent than against SAR-CoV-1 M pro . We determined the crystal structure of ML188 in complex with SARS-CoV-2 M pro to 2.39 Å resolution. Sharing 96% sequence identity, structural comparison of the two complexes only shows subtle differences. Non-covalent protease inhibitors complement the design of covalent inhibitors against SARS-CoV-2 main protease and are critical initial steps in the design of DAAs to treat CoVID 19.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: